Deep learning-based multimodal prediction of chronic kidney disease stage

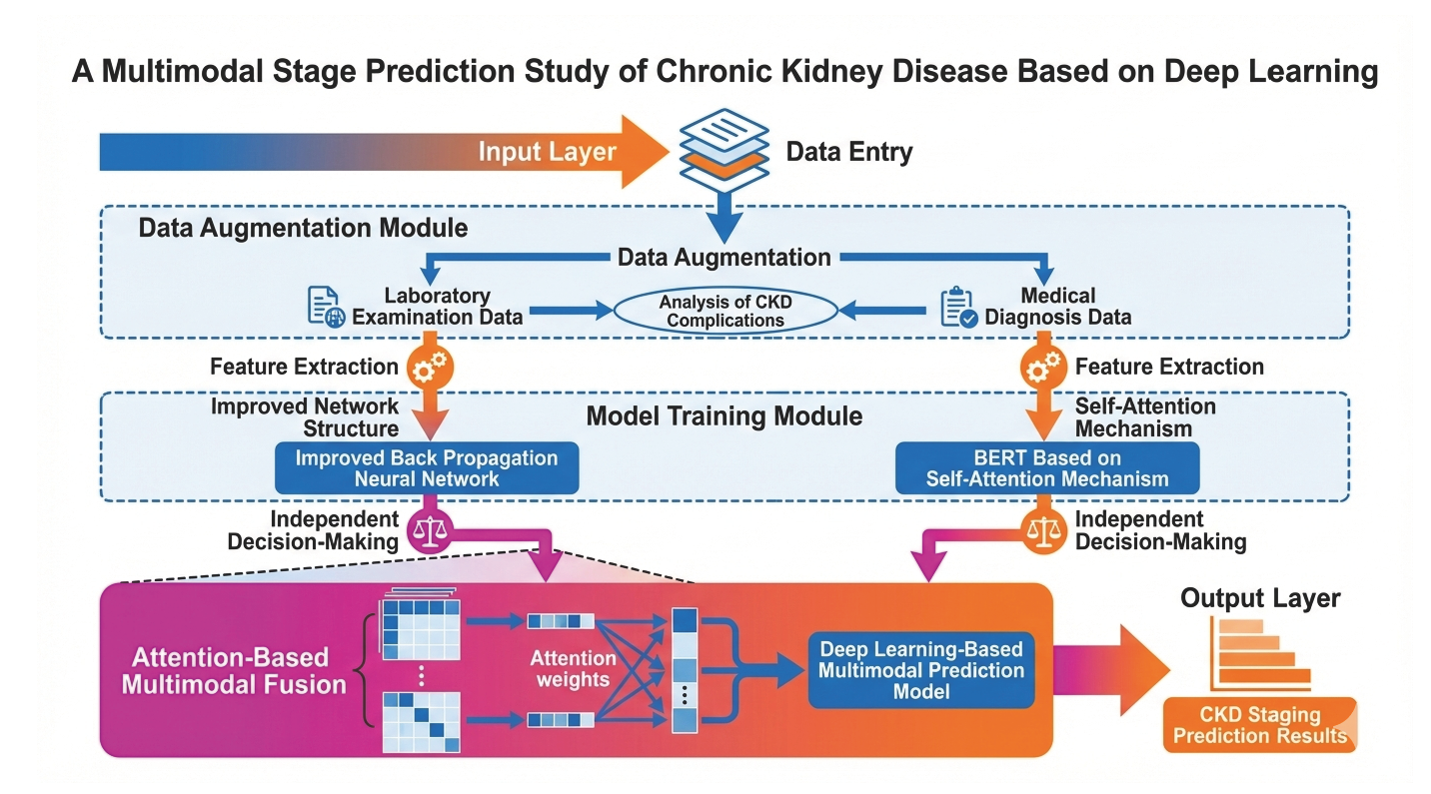
Chronic kidney disease (CKD) constitutes a critical global public health challenge. Its early-stage symptoms are subtle and easily overlooked, frequently resulting in delayed diagnosis and escalated treatment expenditures. To enable timely early intervention, this study proposes a deep learning-based multimodal CKD stage prediction model, which integrates Western medical laboratory data and traditional Chinese medicine (TCM) symptom descriptions, thereby overcoming the inherent limitations of unimodal prediction approaches. For Western medical data, the synthetic minority oversampling technique (SMOTE) was employed to address class imbalance. Subsequently, a least absolute shrinkage and selection operator (LASSO) feature selection model was utilized to identify key biomarkers, including serum creatinine and serum chloride. A backpropagation neural network, enhanced with Adam optimization and regularization mechanisms, was constructed for predictive modeling. For TCM symptom texts (e.g., tongue manifestations and pulse conditions), the Google-pretrained Bidirectional Encoder Representations from Transformers (BERT) model was leveraged to learn latent semantic patterns in the textual data. Finally, a multimodal decision fusion strategy based on the attention mechanism was adopted to dynamically learn the relative importance of Western medical and TCM features in CKD staging prediction, culminating in the development of the proposed deep learning-driven multimodal CKD staging model. Validation results indicate that the proposed model outperforms classical machine learning models and unimodal models across multiple metrics, including accuracy, F1-score, area under the curve, and convergence efficiency. These findings confirm its clinical feasibility and effectiveness, providing an innovative multimodal data-driven prediction framework that synergizes the strengths of Western medical quantitative testing and TCM qualitative diagnosis.
- Zhang L, Wang F, Wang L, et al. Prevalence of chronic kidney disease in China: a cross-sectional survey. Lancet. 2012;379(9818):815-822. doi: 10.1016/S0140-6736(12)60033-6
- Shlipak MG, Tummalapalli SL, Boulware LE, et al. The case for early identification and intervention of chronic kidney disease: conclusions from a Kidney Disease: Improving Global Outcomes (KDIGO) Controversies Conference. Kidney Int. 2021;99(1):34-47. doi: 10.1016/j.kint.2020.10.012
- Hossain MS, Hassa SMN, Al-Amin M, Rahaman MN, Hossain R, Hossain MI. Kidney Disease Detection from CT Images using a customized CNN model and Deep Learning. In: 2023 International Conference on Advances in Intelligent Computing and Applications (AICAPS). IEEE; 2023:1-6. doi: 10.1109/aicaps57044.2023.10074314
- Dubin RF, Deo R, Ren Y, et al. Proteomics of CKD progression in the chronic renal insufficiency cohort. Nat Commun. 2023;14(1):6340. doi: 10.1038/s41467-023-41642-7
- Tin A, Köttgen A. Genome-wide association studies of CKD and related traits. Clin J Am Soc Nephrol. 2020;15(11):1643-1656. doi: 10.2215/cjn.00020120
- Robinson-Cohen C, Triozzi JL, Rowan B, et al. Genomewide association study of CKD progression. J Am Soc Nephrol. 2023;34(9):1547-1559. doi: 10.1681/asn.0000000000000170
- Hannan M, Chen J, Hsu J, et al. Frailty and cardiovascular outcomes in adults with CKD: Findings from the Chronic Renal Insufficiency Cohort (CRIC) study. Am J Kidney Dis. 2024;83(2):208-215. doi: 10.1053/j.ajkd.2023.06.009
- Ding N, Lv Y, Su H, et al. Vascular calcification in CKD: New insights into its mechanisms. J Cell Physiol. 2023;238(6):1160-1182. doi: 10.1002/jcp.31021
- Venkatesan VK, Ramakrishna MT, Izonin I, Tkachenko R, Havryliuk M. Efficient Data Preprocessing with Ensemble Machine Learning Technique for the Early Detection of Chronic Kidney Disease. Appl Sci. 2023;13(5):2885. doi: 10.3390/app13052885
- Krukowski H, Valkenburg S, Madella AM, et al. Gut microbiome studies in CKD: opportunities, pitfalls and therapeutic potential. Nat Rev Nephrol. 2023;19(2):87-101. doi: 10.1038/s41581-022-00647-z
- Kumar K, Pradeepa M, Mahdal M, RajaRao MVLN, Ramesh JVN. A Deep Learning Approach for Kidney Disease Recognition and Prediction through Image Processing. Appl Sci. 2023;13(6):3621. doi: 10.3390/app13063621
- Joshi S, Kalantar-Zadeh K, Chauveau P, Carrero JJ. Risks and Benefits of Different Dietary Patterns in CKD. Am J Kidney Dis. 2023;81(3):352-360. doi: 10.1053/j.ajkd.2022.08.013
- Macdougall IC. Anaemia in CKD—treatment standard. Nephrol Dial Transplant. 2024;39(5):770-777. doi: 10.1093/ndt/gfad250
- Świątek Ł, Jeske J, Miedziaszczyk M, Idasiak-Piechocka I. The impact of a vegetarian diet on chronic kidney disease (CKD) progression–a systematic review. BMC Nephrol. 2023;24(1):168. doi: 10.1186/s12882-023-03233-y
- Yamada S, Nakano T. Role of Chronic Kidney Disease (CKD)–Mineral and Bone Disorder (MBD) in the Pathogenesis of Cardiovascular Disease in CKD. J Atheroscler Thromb. 2023;30(8):835-850. doi: 10.5551/jat.RV22006
- Heo GY, Koh HB, Kim HJ, et al. Association of Plant Protein Intake with Risk of incident CKD: a UK Biobank study. Am J Kidney Dis. 2023;82(6):687-697.e1. doi: 10.1053/j.ajkd.2023.05.007
- Mapuskar KA, Vasquez-Martinez G, Mayoral-Andrade G, Tomanek-Chalkley A, Zepeda-Orozco D, Allen BG. Mitochondrial oxidative metabolism: an emerging therapeutic target to improve CKD outcomes. Biomedicines. 2023;11(6):1573. doi: 10.3390/biomedicines11061573
- Saif D, Sarhan AM, Elshennawy NM. Deep-kidney: an effective deep learning framework for chronic kidney disease prediction. Health Inf Sci Syst. 2023;12(1):3. doi: 10.1007/s13755-023-00261-8
- Alsekait DM, Saleh H, Gabralla LA, et al. Toward comprehensive chronic kidney disease prediction based on ensemble deep learning models. Appl Sci. 2023;13(6):3937. doi: 10.3390/app13063937
- Akter S, Ahmed M, Al Imran A, et al. CKD.Net: A novel deep learning hybrid model for effective, real-time, automated screening tool towards prediction of multi stages of CKD along with eGFR and creatinine. Expert Syst Appl. 2023;223:119851. doi: 10.1016/j.eswa.2023.119851
- Ge XY, Lan ZK, Lan QQ, Lin HS, Wang GD, Chen J. Diagnostic accuracy of ultrasound-based multimodal radiomics modeling for fibrosis detection in chronic kidney disease. Eur Radiol. 2023;33(4):2386-2398. doi: 10.1007/s00330-022-09268-3
- Hou Z, Zhang G, Ma Y, et al. Development of a multimodal kidney age prediction based on automatic segmentation CT image in patients with normal renal function. Clin Kidney J. 2023;16(11):2091-2099. doi: 10.1093/ckj/sfad167
- Chertow GM, Correa-Rotter R, Vart P, et al. Effects of Dapagliflozin in Chronic Kidney Disease, With and Without Other Cardiovascular Medications: DAPA-CKD Trial. J Am Heart Assoc. 2023;12(9):e028739. doi: 10.1161/JAHA.122.028739
- Evenepoel P, Stenvinkel P, Shanahan C. Inflammation and gut dysbiosis as drivers of CKD–MBD. Nat Rev Nephrol. 2023;19(10):646-657. doi: 10.1038/s41581-023-00736-7
- Ren F, Jin Q, Jin Q, et al. Genetic evidence supporting the causal role of gut microbiota in chronic kidney disease and chronic systemic inflammation in CKD: a bilateral twosample Mendelian randomization study. Front Immunol. 2023;14:1287698. doi: 10.3389/fimmu.2023.1287698
- Venkatrao K, Kareemulla S. HDLNET: a hybrid deep learning network model with intelligent IoT for detection and classification of chronic kidney disease. IEEE Access. 2023;11:99638-99652. doi: 10.1109/ACCESS.2023.3312183
- Joo YS, Rim TH, Koh HB, et al. Non-invasive chronic kidney disease risk stratification tool derived from retinabased deep learning and clinical factors. NPJ Digit Med. 2023;6(1):114. doi: 10.1038/s41746-023-00860-5
- Liang P, Yang J, Wang W, et al. Deep learning identifies intelligible predictors of poor prognosis in chronic kidney disease. IEEE J Biomed Health Inform. 2023;27(7):3677-3685. doi: 10.1109/JBHI.2023.3266587
- Yang J, Chen X, Luo C, et al. Application of serum SERS technology combined with deep learning algorithm in the rapid diagnosis of immune diseases and chronic kidney disease. Sci Rep. 2023;13(1):15719. doi: 10.1038/s41598-023-42719-5
- Bhattacharjee A, Rabea S, Bhattacharjee A, et al. A multiclass deep learning model for early lung cancer and chronic kidney disease detection using computed tomography images. Front Oncol. 2023;13:1193746. doi: 10.3389/fonc.2023.1193746
- Zhang M, Ye Z, Yuan E, et al. Imaging-based deep learning in kidney diseases: recent progress and future prospects. Insights Imaging. 2024;15(1):50. doi: 10.1186/s13244-024-01636-5
- Arif MS, Rehman AU, Asif D. Explainable machine learning models for chronic kidney disease prediction. Algorithms. 2024;17(10):443. doi: 10.3390/a17100443
- Arif MS, Mukheimer A, Asif D. Enhancing the Early Detection of Chronic Kidney Disease: A Robust Machine Learning Model. Big Data Cogn Comput. 2023;7(3):144. doi: 10.3390/bdcc7030144
- Li J, Zakka KR, Booth J, et al. Discovering patient groups in sequential electronic healthcare data using unsupervised representation learning. BMC Med Inform Decis Mak. 2025;25(1):45. doi: 10.1186/s12911-024-02812-9
- Miotto R, Li L, Kidd BA, Dudley JT. Deep patient: An unsupervised representation to predict the future of patients from the electronic health records. Sci Rep. 2016;6(1):26094. doi: 10.1038/srep26094
- Friedman SF, Khurshid S, Venn RA, et al. Unsupervised deep learning of electrocardiograms enables scalable human disease profiling. NPJ Digit Med. 2025;8:23. doi: 10.1038/s41746-024-01418-9
- Baltrušaitis T, Ahuja C, Morency LP. Multimodal machine learning: A survey and taxonomy. IEEE Trans Pattern Anal Mach Intell. 2018;41(2):423-443. doi: 10.1109/TPAMI.2018.2798607
- Li W, Ge X, Liu S, Xu L, Zhai X, Yu L. Opportunities and challenges of traditional Chinese medicine doctors in the era of artificial intelligence. Front Med. 2024;10:1336175. doi: 10.3389/fmed.2023.1336175
- Zhai Y, Gu J, Zong F, Jiang W. Fine grained cross-modal retrieval algorithm for IETM with attention mechanism fused. Syst Eng Electron. 2023;45(12):3915-3923. doi: 10.12305/j.issn.1001-506X.2023.12.21
- Chawla NV, Bowyer KW, Hall LO, Kegelmeyer WP. SMOTE: Synthetic minority over-sampling technique. J Artif Intell Res. 2002;16:321-357. doi: 10.1613/jair.953
- Hastie T, Tibshirani R, Friedman J. The Elements of Statistical Learning: Data Mining, Inference, and Prediction. 2nd ed. Springer; 2009. doi: 10.1007/978-0-387-84858-7
- Khatri M, Zitovsky J, Lee D, Nayyar K, Fazzari M, Grant C. The association between serum chloride levels and chronic kidney disease progression: a cohort study. BMC Nephrol. 2020;21(1):165. doi: 10.1186/s12882-020-01828-3
- Kingma D, Ba J. Adam: A method for stochastic optimization. International Conference on Learning Representations (ICLR). arXiv. Preprint posted online January 30, 2017. doi: 10.48550/arXiv.1412.6980
- Bezinque A, Parker J, Babitz SK, Noyes SL, Hu S, Lane BR. Proteinuria impacts patient survival differentially based on clinical setting: A retrospective cross-sectional analysis of cohorts from a single health system: Retrospective cohort study. Ann Med Surg. 2019;45:120-126. doi: 10.1016/j.amsu.2019.07.029
- Feng Y, Shen J, Xu H, Wang R, Liu T, Zhang Y. Cross fusion multimodal sentiment analysis model based on translate mechanism. Data Anal Knowl Discov. 2025;9(3):16-27. doi: 10.11925/infotech.2096-3467.2024.0003
- Bahdanau D, Cho K, Bengio Y. Neural Machine Translation by Jointly Learning to Align and Translate. International Conference on Learning Representations (ICLR). arXiv. Preprint posted online May 19, 2014. doi: 10.48550/arXiv.1409.0473
- Li F, Zheng T, He N, et al. Data-driven hybrid neural fuzzy network and ARX modeling approach to practical industrial process identification. IEEE/CAA J Autom Sin. 2022;9(9):1702-1705. doi: 10.1109/JAS.2022.105821

