Identification and characterization of novel outer membrane proteins of Brachyspira pilosicoli

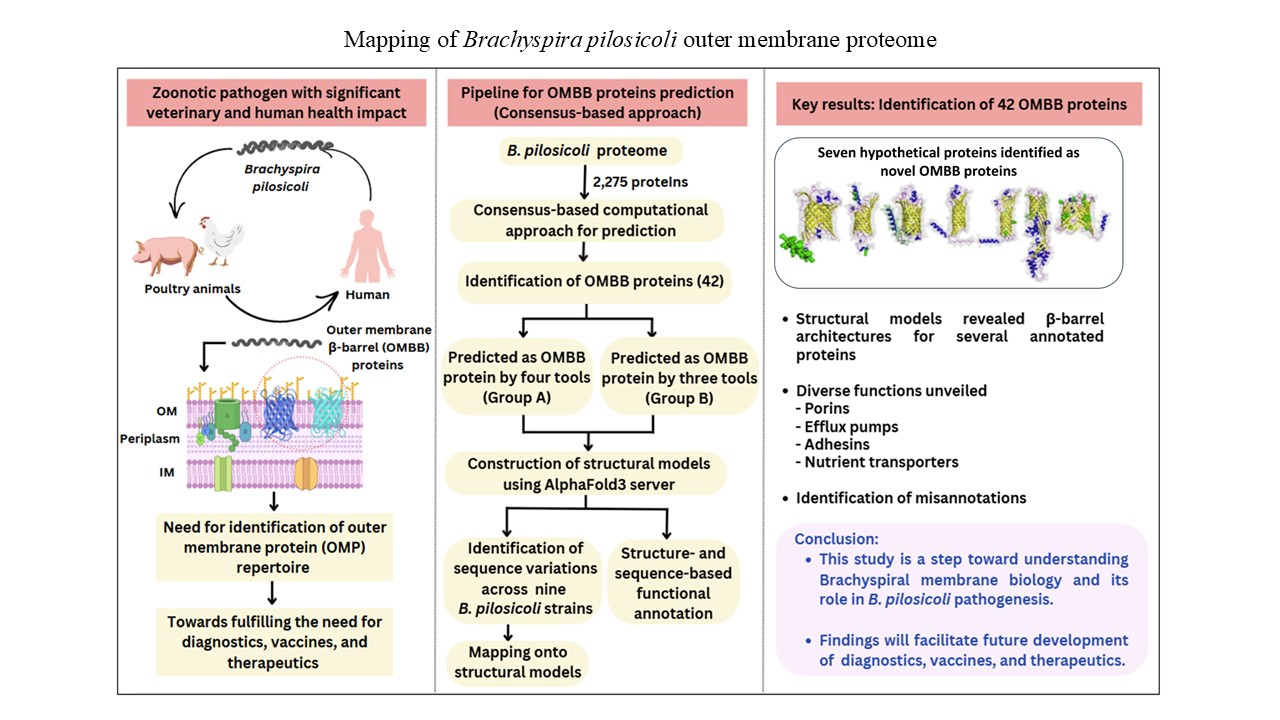
Brachyspira pilosicoli is a globally prevalent, anaerobic, Gram-negative spirochete that causes intestinal spirochetosis in birds, pigs, and humans. It colonizes the large intestine, causing colitis, diarrhea, and impaired growth. Despite its pathogenic relevance, the outer membrane proteins of B. pilosicoli remain largely uncharacterized. In this study, we computationally identified a total of 42 outer membrane β-barrel (OMBB) proteins within the B. pilosicoli proteome using a consensus-based computational framework. Structural models generated using AlphaFold 3 confirmed the β-barrel architectures of the predicted proteins. Structure- and sequence-based functional annotations revealed homologs of b-barrel assembly machinery BamA protein, lipopolysaccharide-assembly protein LPS-assembly protein D, TolC, transporter proteins, enzymes, diffusion channels, and porins. Notably, seven of the predicted OMBB proteins were previously unannotated in UniProt and the National Center for Biotechnology Information; we report their putative functions here for the 1st time. Sequence variation analysis among the homologs of OMBB proteins across nine B. pilosicoli strains revealed that many of the variations were present within surface-exposed loop regions, suggesting roles in host interaction and immune modulation. Our in silico study expands the functional repertoire of B. pilosicoli outer membrane proteins, highlighting potential targets for diagnostics, vaccine development, and therapeutic interventions.
- Ludwig W, Euzéby J, Whitman WB. Taxonomic outlines of the phyla bacteroidetes, spirochaetes, tenericutes (mollicutes), acidobacteria, fibrobacteres, fusobacteria, dictyoglomi, gemmatimonadetes, lentisphaerae, verrucomicrobia, chlamydiae, and planctomycetes. In: Krieg NR, Staley JT, Brown DR, et al, editors. Bergey’s Manual® of Systematic Bacteriology: Volume Four The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes. New York: Springer; 2010. p. 21-24.
- Hampson DJ, Oxberry SL, La T. Potential for zoonotic transmission of Brachyspira pilosicoli. Emerg Infect Dis. 2006;12(5):869-870. doi: 10.3201/eid1205.051180
- Le Roy CI, Mappley LJ, La Ragione RM, Woodward MJ, Claus SP. Brachyspira pilosicoli-induced avian intestinal spirochaetosis. Microb Ecol Health Dis. 2015;26(1):28853. doi: 10.3402/mehd.v26.28853
- Bano L, Merialdi G, Bonilauri P, et al. Prevalence, disease associations and risk factors for colonization with intestinal spirochaetes (Brachyspira spp.) in flocks of laying hens in North-Eastern Italy. Avian Pathol. 2008;37(3):281-286. doi: 10.1080/03079450802043726
- Myers SE, Dunn PA, Phillips ND, La T, Hampson DJ. Brachyspira intermedia and Brachyspira pilosicoli are commonly found in older laying flocks in Pennsylvania. Avian diseases. 2009;53(4):533-537. doi: 10.1637/8900-042709-Reg.1
- Amin MM, Phillips ND, La T, Robertson ID, Hampson DJ. Intestinal spirochaetes (Brachyspira spp.) colonizing flocks of layer and breeder chickens in Malaysia. Avian Pathol. 2014;43(6):501-505. doi: 10.1080/03079457.2014.966056
- Zarabi M, Jamshidi A, Khanzadi S, Razmyar J. Intestinal colonization of different Brachyspira spp. In laying hens. Iran J Vet Med. 2014;8:213-218.
- Hampson DJ. The spirochete Brachyspira pilosicoli, enteric pathogen of animals and humans. Clin Microbiol Rev. 2018;31(1):10.1128/cmr.00087-17. doi: 10.1128/cmr.00087-17
- Smith JL. Colonic spirochetosis in animals and humans. J Food Prot. 2005;68(7):1525-1534. doi: 10.4315/0362-028X-68.7.1525
- Trott DJ, Stanton TB, Jensen NS, Duhamel GE, Johnson JL, Hampson DJ. Serpulina pilosicoli sp. Nov., the agent of porcine intestinal spirochetosis. Int J Syst Bacteriol. 1996;46(1):206-215. doi: 10.1099/00207713-46-1-206
- Stephens CP, Hampson DJ. Intestinal spirochete infections of chickens: A review of disease associations, epidemiology and control. Anim Health Res Rev. 2001;2(1):83-91.
- Gad A, Willén R, Furugård K, Fors B, Hradsky M. Intestinal spirochaetosis as a cause of longstanding diarrhoea. Ups J Med Sci. 1977;82(1):49-54. doi: 10.3109/03009737709179059
- Gan J, Bryant C, Arul D, Parmar C. Intestinal spirochaetosis mimicking acute appendicitis with review of the literature. BMJ Case Rep. 2017;2017:bcr-2017-221574. doi: 10.1136/bcr-2017-221574
- Nishii S, Higashiyama M, Ogata S, et al. Human intestinal spirochetosis mimicking ulcerative colitis. Clin J Gastroenterol. 2018;11:145-149. doi: 10.1007/s12328-017-0807-3
- Marthinsen L, Willén R, Carlén B, Lindberg E, Värendh G. Intestinal spirochetosis in eight pediatric patients from Southern Sweden. APMIS. 2002;110(7‐8):571-579. doi: 10.1034/j.1600-0463.2002.11007809.x
- Oxberry SL, Trott DJ, Hampson DJ. Serpulina pilosicoli, waterbirds and water: Potential sources of infection for humans and other animals. Epidemiol Infect. 1998;121(1):219-225. doi: 10.1017/S0950268898008863
- Trott DJ, Mikosza AS, Combs BG, Oxberry SL, Hampson DJ. Population genetic analysis of Serpulina pilosicoli and its molecular epidemiology in villages in the Eastern Highlands of Papua New Guinea. Int J Syst Bacteriol. 1998;48(3):659-668. doi: 10.1099/00207713-48-3-659
- Hampson DJ, Lugsomya K, La T, Phillips ND, Trott DJ, Abraham S. Antimicrobial resistance in Brachyspira - an increasing problem for disease control. Vet Microbiol. 2019;229:59-71. doi: 10.1016/j.vetmic.2018.12.019
- Christodoulides M, De Oliveira D, Cleary DW, Humbert MV, Machado-De-Ávila RA, La Ragione RM. An in silico reverse vaccinology study of Brachyspira pilosicoli, the causative organism of intestinal spirochaetosis, to identify putative vaccine candidates. Process Biochem. 2022;122:128-148. doi: 10.1016/j.procbio.2022.08.014
- Wanchanthuek P, Bellgard MI, La T, et al. The complete genome sequence of the pathogenic intestinal spirochete Brachyspira pilosicoli and comparison with other Brachyspira genomes. PLoS One. 2010;5(7):e11455. doi: 10.1371/journal.pone.0011455
- Erlandson KM, Klingler ET. Intestinal spirochetosis: Epidemiology, microbiology, and clinical significance. Clin Microbiol Newsl. 2005;27(12):91-96. doi: 10.1016/j.clinmicnews.2005.05.002
- Trott DJ, Alt DP, Zuerner RL, et al. Identification and cloning of the gene encoding BmpC: an outer-membrane lipoprotein associated with Brachyspira pilosicoli membrane vesicles. Microbiology (Reading). 2004;150(4):1041-1053. doi: 10.1099/mic.0.26755-0
- Lee BJ, Hampson DJ. Lipo-oligosaccharide profiles of Serpulina pilosicoli strains and their serological cross-reactivities. J Med Microbiol. 1999;48(4):411-415. doi: 10.1099/00222615-48-4-411
- Koebnik R, Locher KP, Van Gelder P. Structure and function of bacterial outer membrane proteins: Barrels in a nutshell. Mol Microbiol. 2000;37(2):239-253. doi: 10.1046/j.1365-2958.2000.01983.x
- Fairman JW, Noinaj N, Buchanan SK. The structural biology of β-barrel membrane proteins: A summary of recent reports. Curr Opin Struct Biol. 2011;21(4):523-531. doi: 10.1016/j.sbi.2011.05.005
- Horne JE, Brockwell DJ, Radford SE. Role of the lipid bilayer in outer membrane protein folding in gram-negative bacteria. J Biol Chem. 2020;295(30):10340-10367. doi: 10.1074/jbc.REV120.011473
- Kleinschmidt JH. Folding of β-barrel membrane proteins in lipid bilayers - unassisted and assisted folding and insertion. Biochim Biophys Acta. 2015;1848(9):1927-1943. doi: 10.1016/j.bbamem.2015.05.004
- Solan R, Pereira J, Lupas AN, Kolodny R, Ben- Tal N. Gram-negative outer-membrane proteins with multiple β-barrel domains. Proc Natl Acad Sci U S A. 2021;118(31):e2104059118. doi: 10.1073/pnas.2104059118
- Trott DJ, Alt DP, Zuerner RL, Wannemuehler MJ, Stanton TB. The search for Brachyspira outer membrane proteins that interact with the host. Anim Health Res Rev. 2001;2(1):19-30. doi: 10.1079/AHRR200112
- Tenaya IW, Penhale WJ, Hampson DJ. Preparation of diagnostic polyclonal and monoclonal antibodies against outer envelope proteins of Serpulina pilosicoli. J Med Microbiol. 1998;47(4):317-324. doi: 10.1099/00222615-47-4-317
- La T, Phillips ND, Hampson DJ. Vaccination of chickens with the 34 kDa carboxy-terminus of Bpmp72 reduces colonization with Brachyspira pilosicoli following experimental infection. Avian Pathol. 2019;48(1):80-85. doi: 10.1080/03079457.2018.1546377
- Medicine NLo. National Center for Biotechnology Information (NCBI). Bethesda (MD): Medicine NLo; 1988. doi: 10.1093/nar/gkaa892
- Rice P, Longden I, Bleasby A. EMBOSS: The European molecular biology open software suite. Trends Genet. 2000;16(6):276-277. doi: 10.1016/S0168-9525(00)02024-2
- Teufel F, Almagro Armenteros JJ, Johansen AR, et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat Biotechnol. 2022;40(7):1023-1025. doi: 10.1038/s41587-021-01156-3
- Juncker AS, Willenbrock H, Von Heijne G, Brunak S, Nielsen H, Krogh A. Prediction of lipoprotein signal peptides in gram-negative bacteria. Protein Sci. 2003;12(8):1652-1662. doi: 10.1110/ps.0303703
- Yu CS, Lin CJ, Hwang JK. Predicting subcellular localization of proteins for gram-negative bacteria by support vector machines based on n-peptide compositions. Protein Sci. 2004;13(5):1402-1406. doi: 10.1110/ps.03479604
- Yu NY, Wagner JR, Laird MR, et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics. 2010;26(13):1608-1615. doi: 10.1093/bioinformatics/btq249
- Luo H, Lin Y, Liu T, et al. DEG 15, an update of the database of essential genes that includes built-in analysis tools. Nucleic Acids Res. 2021;49(D1):D677-D686. doi: 10.1093/nar/gkaa917
- Roumia AF, Tsirigos KD, Theodoropoulou MC, Tamposis IA, Hamodrakas SJ, Bagos PG. OMPdb: A global hub of beta-barrel outer membrane proteins. Front Bioinform. 2021;1:646581. doi: 10.3389/fbinf.2021.646581
- Bagos PG, LiakopoulipopTD, Hamodrakas SJ. Finding beta-barrel outer membrane proteins with a markov chain model. WSEAS Trans Biol Biomed. 2004;2(1):186-189.
- Ou YY, Gromiha MM, Chen SA, Suwa M. TMBETADISC-RBF: Discrimination of beta-barrel membrane proteins using RBF networks and PSSM profiles. Comput Biol Chem. 2008;32(3):227-231. doi: 10.1016/j.compbiolchem.2008.03.002
- Bernhofer M, Rost B. TMbed: Transmembrane proteins predicted through language model embeddings. BMC Bioinformatics. 2022;23(1):326. doi: 10.1186/s12859-022-04873-x
- Abramson J, Adler J, Dunger J, et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature. 2024;630:493-500. doi: 10.1038/s41586-024-07487-w
- Xu J, Zhang Y. How significant is a protein structure similarity with TM-score= 0.5? Bioinformatics. 2010;26(7):889-895. doi: 10.1093/bioinformatics/btq066
- Zhang Y, Skolnick J. Scoring function for automated assessment of protein structure template quality. Proteins. 2004;57(4):702-710. doi: 10.1002/prot.20264
- DeLano WL. The PyMOL Molecular Graphics System; 2002. https://www/pymol/org [Last accessed on 2025 Jan 25].
- Lin Z, Akin H, Rao R, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science. 2023;379(6637):1123-1130. doi: 10.1126/science.ade2574
- Waterhouse A, Bertoni M, Bienert S, et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018;46(W1):W296-W303. doi: 10.1093/nar/gky427
- Baek M, DiMaio F, Anishchenko I, et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science. 2021;373(6557):871-876. doi: 10.1126/science.abj8754
- Du Z, Su H, Wang W, et al. The trRosetta server for fast and accurate protein structure prediction. Nat Protoc. 2021;16(12):5634-5651. doi: 10.1038/s41596-021-00628-9
- Holm L. Dali server: Structural unification of protein families. Nucleic Acids Res. 2022;50(W1):W210-W215. doi: 10.1093/nar/gkac387
- Holm L. Using dali for protein structure comparison. Methods Mol Biol. 2020;2112:29-42. doi: 10.1007/978-1-0716-0270-6_3
- Van Kempen M, Kim SS, Tumescheit C, et al. Fast and accurate protein structure search with foldseek. Nat Biotechnol. 2024;42(2):243-246. doi: 10.1038/s41587-023-01773-0
- Törönen P, Holm L. PANNZER-a practical tool for protein function prediction. Protein Sci. 2022;31(1):118-128. doi: 10.1002/pro.4193
- Cantalapiedra CP, Hernández-Plaza A, Letunic I, Bork P, Huerta-Cepas J. EggNOG-mapper v2: Functional annotation, orthology assignments, and domain prediction at the metagenomic scale. Mol Biol Evol. 2021;38(12):5825-5829. doi: 10.1093/molbev/msab293
- Larkin MA, Blackshields G, Brown NP, et al. Clustal W and clustal X version 2.0. Bioinformatics. 2007;23(21):2947-2948. doi: 10.1093/bioinformatics/btm404
- Kumar S, Stecher G, Suleski M, Sanderford M, Sharma S, Tamura K. MEGA12: Molecular evolutionary genetic analysis version 12 for adaptive and green computing. Mol Biol Evol. 2024;41(12):msae263. doi: 10.1093/molbev/msae263
- Zhang C, Shine M, Pyle AM, Zhang Y. US-align: Universal structure alignments of proteins, nucleic acids, and macromolecular complexes. Nat Methods. 2022;19(9):1109-1115. doi: 10.1038/s41592-022-01585-1
- Derbyshire MC. Bioinformatic detection of positive selection pressure in plant pathogens: The neutral theory of molecular sequence evolution in action. Front Microbiol. 2020;11:644. doi: 10.3389/fmicb.2020.00644
- Bakelar J, Buchanan SK, Noinaj N. The structure of the β-barrel assembly machinery complex. Science. 2016;351(6269):180-186. doi: 10.1126/science.aad3460
- Konovalova A, Kahne DE, Silhavy TJ. Outer membrane biogenesis. Annu Rev Microbiol. 2017;71(1):539-556. doi: 10.1146/annurev-micro-090816-093754
- Lysnyansky I, Ron Y, Sachse K, Yogev D. Intrachromosomal recombination within the vsp locus of Mycoplasma bovis generates a chimeric variable surface lipoprotein antigen. Infect Immun. 2001;69(6):3703-3712. doi: 10.1128/iai.69.6.3703-3712.2001
- Gabe JD, Dragon E, Chang RJ, McCaman MT. Identification of a linked set of genes in Serpulina hyodysenteriae (B204) predicted to encode closely related 39-kilodalton extracytoplasmic proteins. J Bacteriol. 1998;180(2):444-448. doi: 10.1128/jb.180.2.444-448.1998
- Gömmel M, Barth S, Heydel C, Baljer G, Herbst W. Adherence of Brachyspira hyodysenteriae to porcine intestinal epithelial cells is inhibited by antibodies against outer membrane proteins. Curr Microbiol. 2013;66:286-292. doi: 10.1007/s00284-012-0267-4
- Bellgard MI, Wanchanthuek P, La T, et al. Genome sequence of the pathogenic intestinal spirochete Brachyspira hyodysenteriae reveals adaptations to its lifestyle in the porcine large intestine. PLoS One. 2009;4(3):e4641. doi: 10.1371/journal.pone.0004641
- Wang Y, Pannuri AA, Ni D, et al. Structural basis for translocation of a biofilm-supporting exopolysaccharide across the bacterial outer membrane. J Biol Chem. 2016;291(19):10046-10057. doi: 10.1074/jbc.M115.711762
- Itoh Y, Rice JD, Goller C, et al. Roles of pgaABCD genes in synthesis, modification, and export of the Escherichia coli biofilm adhesin poly-beta-1,6-N-acetyl-D-glucosamine. J Bacteriol. 2008;190(10):3670-3680. doi: 10.1128/jb.01920-07
- Wang X, Preston JF 3rd, Romeo T. The pgaABCD locus of Escherichia coli promotes the synthesis of a polysaccharide adhesin required for biofilm formation. J Bacteriol. 2004;186(9):2724-2734. doi: 10.1128/jb.186.9.2724-2734.2004
- Bellgard M, Hampson DJ, La T. Genes and Proteins of Brachyspira Hyodysenteriae and uses Thereof. Google Patents; 2015. 70. Zimmer J. A molecular description of cellulose biosynthesis. Biophys J. 2015;108(2):499a. doi: 10.1016/j.bpj.2014.11.2734
- Römling U, Galperin MY. Bacterial cellulose biosynthesis: Diversity of operons, subunits, products, and functions. Trends Microbiol. 2015;23(9):545-557. doi: 10.1016/j.tim.2015.05.005
- Acheson JF, Derewenda ZS, Zimmer J. Architecture of the cellulose synthase outer membrane channel and its association with the periplasmic TPR domain. Structure. 2019;27(12):1855-1861.e3. doi: 10.1016/j.str.2019.09.008
- Mathes A, Engelhardt H. Nonlinear and asymmetric open channel characteristics of an ion-selective porin in planar membranes. Biophys J. 1998;75(3):1255-1262. doi: 10.1016/S0006-3495(98)74045-7
- Zeth K, Diederichs K, Welte W, Engelhardt H. Crystal structure of Omp32, the anion-selective porin from comamonas acidovorans, in complex with a periplasmic peptide at 2.1 Å resolution. Structure. 2000;8(9):981-992. doi: 10.1016/S0969-2126(00)00189-1
- Azghani AO, Idell S, Bains M, Hancock RE. Pseudomonas aeruginosa outer membrane protein F is an adhesin in bacterial binding to lung epithelial cells in culture. Microb Pathog. 2002;33(3):109-114. doi: 10.1006/mpat.2002.0514
- Wu L, Estrada O, Zaborina O, et al. Recognition of host immune activation by Pseudomonas aeruginosa. Science. 2005;309(5735):774-777. doi: 10.1126/science.1112422
- Fito-Boncompte L, Chapalain A, Bouffartigues E, et al. Full virulence of Pseudomonas aeruginosa requires OprF. Infect Immunity. 2011;79(3):1176-1186. doi: 10.1128/iai.00850-10
- Wessel AK, Liew J, Kwon T, Marcotte EM, Whiteley M. Role of Pseudomonas aeruginosa peptidoglycan-associated outer membrane proteins in vesicle formation. J Bacteriol. 2013;195(2):213-219. doi: 10.1128/jb.01253-12
- Abellón-Ruiz J, Zahn M, Baslé A, Van Den Berg B. Crystal structure of the Acinetobacter baumannii outer membrane protein Omp33. Acta Crystallogr D Struct Biol. 2018;74(9):852-860. doi: 10.1107/S205979831800904X
- Alhede M, Bjarnsholt T, Givskov M, Alhede M. Pseudomonas aeruginosa biofilms: Mechanisms of immune evasion. Adv Appl Microbiol. 2014;86:1-40. doi: 10.1016/B978-0-12-800262-9.00001-9
- Mayeux G, Gayet L, Liguori L, et al. Cell-free expression of the outer membrane protein OprF of Pseudomonas aeruginosa for vaccine purposes. Life Sci Alliance. 2021;4(6):e202000958. doi: 10.26508/lsa.202000958
- Goyal P, Krasteva PV, Van Gerven N, et al. Structural and mechanistic insights into the bacterial amyloid secretion channel CsgG. Nature. 2014;516(7530):250-253. doi: 10.1038/nature13768
- La Ragione RM, Sayers AR, Woodward MJ. The role of fimbriae and flagella in the colonization, invasion and persistence of Escherichia coli O78:K80 in the day-old-chick model. Epidemiol Infect. 2000;124(3):351-363. doi: 10.1017/S0950268899004045
- La Ragione RM, Cooley WA, Woodward MJ. The role of fimbriae and flagella in the adherence of avian strains of Escherichia coli O78:K80 to tissue culture cells and tracheal and gut explants. J Med Microbiol. 2000;49(4):327-338. doi: 10.1099/0022-1317-49-4-327
- La Ragione RM, Woodward MJ. Virulence factors of Escherichia coli serotypes associated with avian colisepticaemia. Res Vet Sci. 2002;73(1):27-35. doi: 10.1016/S0034-5288(02)00075-9
- Vandeputte-Rutten L, Bos MP, Tommassen J, Gros P. Crystal structure of neisserial surface protein A (NspA), a conserved outer membrane protein with vaccine potential. J Biol Chem. 2003;278(27):24825-24830. doi: 10.1074/jbc.M302803200
- Martin D, Cadieux N, Hamel J, Brodeur BR. Highly conserved Neisseria meningitidis surface protein confers protection against experimental infection. J Exp Med. 1997;185(7):1173-1184. doi: 10.1084/jem.185.7.1173
- Moe GR, Tan S, Granoff DM. Differences in surface expression of NspA among Neisseria meningitidis group B strains. Infect Immun. 1999;67(11):5664-5675. doi: 10.1128/iai.67.11.5664-5675.1999
- Moe GR, Zuno-Mitchell P, Lee SS, Lucas AH, Granoff DM. Functional activity of anti-neisserial surface protein A monoclonal antibodies against strains of Neisseria meningitidis serogroup B. Infect Immun. 2001;69(6):3762-3771. doi: 10.1128/iai.69.6.3762-3771.2001
- Koronakis V, Eswaran J, Hughes C. Structure and function of TolC: The bacterial exit duct for proteins and drugs. Annu Rev Biochem. 2004;73(1):467-489. doi: 10.1146/annurev.biochem.73.011303.074104
- Koronakis V, Sharff A, Koronakis E, Luisi B, Hughes C. Crystal structure of the bacterial membrane protein TolC central to multidrug efflux and protein export. Nature. 2000;405(6789):914-919. doi: 10.1038/35016007
- Vakharia H, German GJ, Misra R. Isolation and characterization of Escherichia coli tolC mutants defective in secreting enzymatically active alpha-hemolysin. J Bacteriol. 2001;183(23):6908-6916. doi: 10.1128/jb.183.23.6908-6916.2001
- Wandersman C, Delepelaire P. TolC, an Escherichia coli outer membrane protein required for hemolysin secretion. Proc Natl Acad Sci. 1990;87(12):4776-4780. doi: 10.1073/pnas.87.12.4776
- Clowes R. Transmission and elimination of colicin factors and some aspects of immunity to colicin E1 in Escherichia coli. Zentrabl Bakteriol Parasiten Abt I Orig. 1965;196:152-157.
- Davies JK, Reeves P. Genetics of resistance to colicins in Escherichia coli K-12: Cross-resistance among colicins of group A. J Bacteriol. 1975;123(1):102-117. doi: 10.1128/jb.123.1.102-117.1975
- Fralick JA. Evidence that TolC is required for functioning of the Mar/AcrAB efflux pump of Escherichia coli. J Bacteriol. 1996;178(19):5803-5805. doi: 10.1128/jb.178.19.5803-5805.1996
- German GJ, Misra R. The TolC protein of Escherichia coli serves as a cell-surface receptor for the newly characterized TLS bacteriophage. J Mol Biol. 2001;308(4):579-585. doi: 10.1006/jmbi.2001.4578
- Fredriksson R, Lagerström MC, Lundin LG, Schiöth HB. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol Pharmacol. 2003;63(6):1256-1272. doi: 10.1124/mol.63.6.1256
- Pierce KL, Premont RT, Lefkowitz RJ. Seven-transmembrane receptors. Nat Rev Mol Cell Biol. 2002;3(9):639-650. doi: 10.1038/nrm908
- Consortium TU. UniProt: The universal protein knowledgebase in 2025. Nucleic Acids Res. 2024;53(D1):D609-D617. doi: 10.1093/nar/gkae1010
- Van Alphen L, Havekes L, Lugtenberg B. Major outer membrane protein d of Escherichia coli K12. Purification and in vitro activity of bacteriophage k3 and f-pilus mediated conjugation. FEBS Lett. 1977;75:285-290. doi: 10.1016/0014-5793(77)80104-x
- Datta DB, Arden B, Henning U. Major proteins of the Escherichia coli outer cell envelope membrane as bacteriophage receptors. J Bacteriol. 1977;131(3):821-829. doi: 10.1128/jb.131.3.821-829.1977
- Chai T, Foulds J. Demonstration of a missing outer membrane protein in tolG mutants of Escherichia cell. J Mol Biol. 1974;85(3):465-474. doi: 10.1016/0022-2836(74)90445-8
- Skurray RA, Hancock RE, Reeves P. Con--mutants: Class of mutants in Escherichia coli K-12 lacking a major cell wall protein and defective in conjugation and adsorption of a bacteriophage. J Bacteriol. 1974;119(3):726-735. doi: 10.1128/jb.119.3.726-735.1974
- Schweizer M, Henning U. Action of a major outer cell envelope membrane protein in conjugation of Escherichia coli K-12. J Bacteriol. 1977;129(3):1651-1652. doi: 10.1128/jb.129.3.1651-1652.1977
- Braun V. Covalent lipoprotein from the outer membrane of Escherichia coli. Biochim Biophys Acta. 1975;415(3):335-377. doi: 10.1016/0304-4157(75)90013-1
- Sonntag I, Schwarz H, Hirota Y, Henning U. Cell envelope and shape of Escherichia coli: Multiple mutants missing the outer membrane lipoprotein and other major outer membrane proteins. J Bacteriol. 1978;136(1):280-285. doi: 10.1128/jb.136.1.280-285.1978
- Sugawara E, Nikaido H. Pore-forming activity of OmpA protein of Escherichia coli. J Biol Chem. 1992;267(4):2507-2511. doi: 10.1016/S0021-9258(18)45908-X
- Krishnan S, Prasadarao NV. Outer membrane protein A and OprF: Versatile roles in gram-negative bacterial infections. FEBS J. 2012;279(6):919-931. doi: 10.1111/j.1742-4658.2012.08482.x
- Laguri C, Sperandeo P, Pounot K, et al. Interaction of lipopolysaccharides at intermolecular sites of the periplasmic Lpt transport assembly. Sci Rep. 2017;7(1):9715. doi: 10.1038/s41598-017-10136-0
- Pieńko T, Trylska J. Extracellular loops of BtuB facilitate transport of vitamin B12 through the outer membrane of E. Coli. PLoS Comput Biol. 2020;16(7):e1008024. doi: 10.1371/journal.pcbi.1008024
- Di Girolamo PM, Kadner RJ, Bradbeer C. Isolation of vitamin B 12 transport mutants of Escherichia coli. J Bacteriol. 1971;106(3):751-757. doi: 10.1128/jb.106.3.751-757.1971
- White JC, Di Girolamo PM, Fu ML, Preston YA, Bradbeer C. Transport of vitamin B 12 in Escherichia coli. Location and properties of the initial B 12 -binding site. J Biol Chem. 1973;248(11):3978-3986. doi: 10.1016/S0021-9258(19)43828-3
- Noinaj N, Guillier M, Barnard TJ, Buchanan SK. TonB-dependent transporters: Regulation, structure, and function. Annu Rev Microbiol. 2010;64(1):43-60. doi: 10.1146/annurev.micro.112408.134247
- Schauer K, Rodionov DA, De Reuse H. New substrates for TonB-dependent transport: Do we only see the ‘tip of the iceberg’? Trends Biochem Sci. 2008;33(7):330-338. doi: 10.1016/j.tibs.2008.04.012
- Biswas S, Mohammad MM, Patel DR, Movileanu L, Van Den Berg B. Structural insight into OprD substrate specificity. Nat Struct Mol Biol. 2007;14(11):1108-1109. doi: 10.1038/nsmb1304
- Moraes TF, Bains M, Hancock RE, Strynadka NC. An arginine ladder in OprP mediates phosphate-specific transfer across the outer membrane. Nat Struct Mol Biol. 2007;14(1):85-87. doi: 10.1038/nsmb1189
- Eren E, Vijayaraghavan J, Liu J, et al. Substrate specificity within a family of outer membrane carboxylate channels. PLoS Biol. 2012;10(1):e1001242. doi: 10.1371/journal.pbio.1001242
- Biswas S, Mohammad MM, Movileanu L, Van Den Berg B. Crystal structure of the outer membrane protein OpdK from Pseudomonas aeruginosa. Structure. 2008;16(7):1027-1035. doi: 10.1016/j.str.2008.04.009
- Kuehne SA, Cartman ST, Heap JT, Kelly ML, Cockayne A, Minton NP. The role of toxin A and toxin B in Clostridium difficile infection. Nature. 2010;467(7316):711-713. doi: 10.1038/nature09397
- Van Den Berg B, Prathyusha Bhamidimarri S, Dahyabhai Prajapati J, Kleinekathöfer U, Winterhalter M. Outer-membrane translocation of bulky small molecules by passive diffusion. Proc Natl Acad Sci. 2015;112(23):E2991-E2999. doi: 10.1073/pnas.1424835112
- Pajatsch M, Andersen C, Mathes A, Böck A, Benz R, Engelhardt H. Properties of a cyclodextrin-specific, unusual porin from Klebsiella oxytoca. J Biol Chem. 1999;274(35):25159-25166. doi: 10.1074/jbc.274.35.25159
- Gruss F, Zähringer F, Jakob RP, Burmann BM, Hiller S, Maier T. The structural basis of autotransporter translocation by TamA. Nat Struct Mol Biol. 2013;20(11):1318-1320. doi: 10.1038/nsmb.2689
- Selkrig J, Mosbahi K, Webb CT, et al. Discovery of an archetypal protein transport system in bacterial outer membranes. Nat Struct Mol Biol. 2012;19(5):506-510. doi: 10.1038/nsmb.2261
- Pavlova O, Peterson JH, Ieva R, Bernstein HD. Mechanistic link between β barrel assembly and the initiation of autotransporter secretion. Proc Natl Acad Sci. 2013;110(10):E938-E947. doi: 10.1073/pnas.1219076110
- Matthijs S, Koedam N, Cornelis P, De Greve H. The trehalose operon of Pseudomonas fluorescens ATCC 17400. Res Microbiol. 2000;151(10):845-851. doi: 10.1016/S0923-2508(00)01151-7
- Abergel C, Bouveret E, Claverie JM, et al. Structure of the Escherichia coli TolB protein determined by MAD methods at 1.95 Å resolution. Structure. 1999;7(10):1291-1300. doi: 10.1016/S0969-2126(00)80062-3
- Rigal A, Bouveret E, Lloubes R, Lazdunski C, Benedetti H. The TolB protein interacts with the porins of Escherichia coli. J Bacteriol. 1997;179(23):7274-7279. doi: 10.1128/jb.179.23.7274-7279.1997
- Fognini-Lefebvre N, Lazzaroni JC, Portalier R. tolA, tolB and excC, three cistrons involved in the control of pleiotropic release of periplasmic proteins by Escherichia coli K12. Mol Gen Genet. 1987;209:391-395. doi: 10.1007/BF00329670
- Lazzaroni JC, Fognini-Lefebvre N, Portalier R. Cloning of the excC and excD genes involved in the release of periplasmic proteins by Escherichia coli K12. Mol Gen Genet. 1989;218:460-464. doi: 10.1007/BF00332410
- Maier T, Clantin B, Gruss F, et al. Conserved Omp85 lid-lock structure and substrate recognition in FhaC. Nat Commun. 2015;6(1):7452. doi: 10.1038/ncomms8452
- Delattre AS, Saint N, Clantin B, et al. Substrate recognition by the POTRA domains of TpsB transporter FhaC. Mol Microbiol. 2011;81(1):99-112. doi: 10.1111/j.1365-2958.2011.07680.x
- Lauber F, Deme JC, Lea SM, Berks BC. Type 9 secretion system structures reveal a new protein transport mechanism. Nature. 2018;564(7734):77-82. doi: 10.1038/s41586-018-0693-y
- Van Den Berg B, Black PN, Clemons WM Jr., Rapoport TA. Crystal structure of the long-chain fatty acid transporter FadL. Science. 2004;304(5676):1506-1509. doi: 10.1126/science.1097524
- Black PN, Said B, Ghosn CR, Beach JV, Nunn WD. Purification and characterization of an outer membrane-bound protein involved in long-chain fatty acid transport in Escherichia coli. J Biol Chem. 1987;262(3):1412-1419. doi: 10.1016/S0021-9258(19)75801-3
- Lauber F, Deme JC, Liu X, et al. Structural insights into the mechanism of protein transport by the type 9 secretion system translocon. Nat Microbiol. 2024;9(4):1089-1102. doi: 10.1038/s41564-024-01644-7
- Otto BR, Sijbrandi R, Luirink J, et al. Crystal structure of hemoglobin protease, a heme binding autotransporter protein from pathogenic Escherichia coli. J Biol Chem. 2005;280(17):17339-17345. doi: 10.1074/jbc.M412885200
- Otto BR, Van Dooren SJ, Dozois CM, Luirink J, Oudega B. Escherichia coli hemoglobin protease autotransporter contributes to synergistic abscess formation and heme-dependent growth of Bacteroides fragilis. Infect Immun. 2002;70(1):5-10. doi: 10.1128/iai.70.1.5-10.2002
- Otto BR, Van Dooren SJ, Nuijens JH, Luirink J, Oudega B. Characterization of a hemoglobin protease secreted by the pathogenic Escherichia coli strain EB1. J Exp Med. 1998;188(6):1091-1103. doi: 10.1084/jem.188.6.1091
- Berg BVD, Bhamidimarri SP, Winterhalter M. Crystal structure of a COG4313 outer membrane channel. Sci Rep. 2015;5(1):11927. doi: 10.1038/srep11927
- Henderson IR, Nataro JP. Virulence functions of autotransporter proteins. Infect Immun. 2001;69(3):1231-1243. doi: 10.1128/iai.69.3.1231-1243.2001
- Yen MR, Peabody CR, Partovi SM, Zhai Y, Tseng YH, Saier MH Jr. Protein-translocating outer membrane porins of gram-negative bacteria. Biochim Biophys Acta. 2002;1562(1-2):6-31. doi: 10.1016/S0005-2736(02)00359-0
- Henderson IR, Navarro-Garcia F, Nataro JP. The great escape: Structure and function of the autotransporter proteins. Trends Microbiol. 1998;6(9):370-378. doi: 10.1016/S0966-842X(98)01318-3
- Dutta PR, Cappello R, Navarro-García F, Nataro JP. Functional comparison of serine protease autotransporters of Enterobacteriaceae. Infect Immun. 2002;70(12):7105-7113. doi: 10.1128/iai.70.12.7105-7113.2002
- Brunder W, Schmidt H, Karch H. EspP, a novel extracellular serine protease of enterohaemorrhagic Escherichia coli O157:H7 cleaves human coagulation factor V. Mol Microbiol. 1997;24(4):767-778. doi: 10.1046/j.1365-2958.1997.3871751.x
- Scandella CJ, Kornberg A. Membrane-bound phospholipase A1 purified from Escherichia coli. Biochemistry. 1971;10(24):4447-4456. doi: 10.1021/bi00800a015
- Pugsley AP, Schwartz M. Colicin E2 release: Lysis, leakage or secretion? Possible role of a phospholipase. EMBO J. 1984;3(10):2393-2397. doi: 10.1002/j.1460-2075.1984.tb02145.x
- Van Der Wal FJ, Luirink J, Oudega B. Bacteriocin release proteins: Mode of action, structure, and biotechnological application. FEMS Microbiol Rev. 1995;17(4):381-399. doi: 10.1111/j.1574-6976.1995.tb00221.x
- De Vries FP, Cole R, Dankert J, Frosch M, Van Putten JP. Neisseria meningitidis producing the Opc adhesin binds epithelial cell proteoglycan receptors. Mol Microbiol. 1998;27(6):1203-1212. doi: 10.1046/j.1365-2958.1998.00763.x
- Virji M, Makepeace K, Moxon ER. Distinct mechanisms of interactions of Opc-expressing meningococci at apical and basolateral surfaces of human endothelial cells; the role of integrins in apical interactions. Mol Microbiol. 1994;14(1):173-184. doi: 10.1111/j.1365-2958.1994.tb01277.x
- Mammarappallil JG, Elsinghorst EA. Epithelial cell adherence mediated by the enterotoxigenic Escherichia coli Tia protein. Infect Immun. 2000;68(12):6595-6601. doi: 10.1128/iai.68.12.6595-6601.2000
- Fleckenstein JM, Kopecko DJ, Warren RL, Elsinghorst EA. Molecular characterization of the tia invasion locus from enterotoxigenic Escherichia coli. Infect Immun. 1996;64(6):2256-2265. doi: 10.1128/iai.64.6.2256-2265.1996
- Gensberg K, Smith AW, Brinkman FS, Hancock RE. Identification of oprG, a gene encoding a major outer membrane protein of Pseudomonas aeruginosa. J Antimicrob Chemother. 1999;43(4):607-608. doi: 10.1093/jac/43.4.607
- Celia H, Noinaj N, Zakharov SD, et al. Structural insight into the role of the ton complex in energy transduction. Nature. 2016;538(7623):60-65. doi: 10.1038/nature19757

