Integrated artificial intelligence frameworks in single-cell multiomics: From intelligent automation to generative modeling

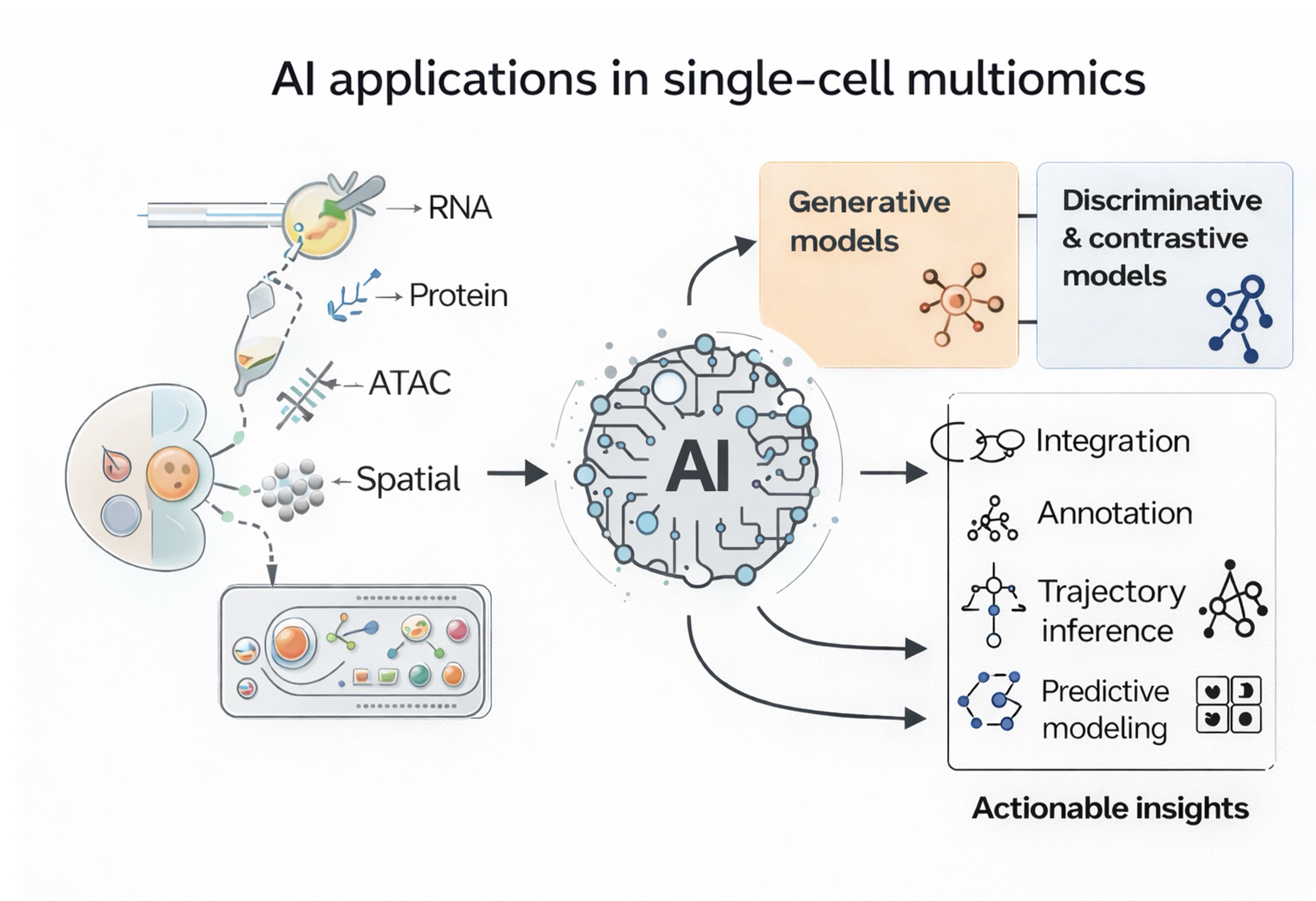
Single-cell multiomics has transformed biomedical research by enabling the study of gene expression, epigenetic modifications, and protein profiles within individual cells at an unprecedented level of detail. This approach has opened new opportunities for understanding complex biological systems, although significant challenges remain. As datasets increase in size and incorporate multiple modalities, issues such as data sparsity, integration complexity, and the need for scalable experimental methods have become prominent. In this review, recent progress combining computational tools, microfluidic technologies, and clinical applications is examined. The first section focuses on how advanced algorithms have been applied to interpret multimodal data, improve cell type identification, map developmental trajectories, and integrate diverse datasets. New techniques incorporating deep learning architectures, such as variational autoencoders, graph neural networks, and emerging foundation models, are highlighted for their role in enabling robust multimodal integration and predictive analysis. Subsequent sections address innovations in experimental workflows. Microfluidic devices integrated with smart automation and real-time monitoring have improved the reliability and efficiency of single-cell studies. These technical advances have had a tangible impact on translational research. In oncology, immunology, and infectious disease, multiomics-driven insights are informing diagnostic strategies and guiding therapeutic development. Finally, remaining challenges are considered, including regulatory requirements and the incorporation of emerging technologies such as spatial omics. Collectively, these advances point toward a future in which single-cell analysis becomes a cornerstone of precision medicine.
- Aswini R, Saranya B, Gayathri K, Karthikeyan E. Revolutionizing infectious disease surveillance: Multiomics technologies and AI-driven integration. Decoding Infect Transm. 2025;3:100061. doi: 10.1016/j.dcit.2025.100061
- Chen TY, You L, Hardillo JA, Chien MP. Spatial transcriptomic technologies. Cells. 2023;12(16):2042. doi: 10.3390/cells12162042
- Lee Y, Lee M, Shin Y, Kim K, Kim T. Spatial Omics in Clinical Research: A Comprehensive Review of Technologies an Guidelines for Applications. Int J Mol Sci. 2025;26(9):3949. doi: 10.3390/ijms26093949
- Hira MT, Razzaque MA, Angione C, Scrivens J, Sawan S, Sarker M. Integrated multi-omics analysis of ovarian cancer using variational autoencoders. Sci Rep. 2021;11(1):6265. doi: 10.1038/s41598-021-85285-4
- Liu D, Chen H, Wang J, Wang Y. MultiGNN: A Graph Neural Network Framework for Inferring Gene Regulatory Networks from Single-Cell Multi-Omics Data. Computation. 2025;13(5):124. doi: 10.3390/computation13050124
- Zhang J, Xue J, Luo N, Chen F, Chen B, Zhao Y. Microwell array chip-based single-cell analysis. Lab Chip. 2023;23(5):1066-1079. doi: 10.1039/D2LC00667G
- Nguyen R, Hlathu Z, Gordon C, et al. Flexible and high-throughput approach for capturing large number of single cells simultaneously using microwell-based technology. J Immunol. 2023;210(Supp 1):251.08. doi: 10.4049/jimmunol.210.Supp.251.08
- Song HW, Acob RA, Hlathu Z, et al. Abstract 333: Stability and performance of a partially used 8-lane microwell cartridge on single-cell capture and immune response profiling. Cancer Res. 2024;84(Supp 6):333. doi: 10.1158/1538-7445.am2024-333
- Zhao X, Chen Y, Jiang Y, Radenbaugh A, Moskwa J, Jensen D. Machine learning approaches for defect detection in a microwell-based medical device. arXiv. Preprint posted online 2024. doi: 10.48550/arXiv.2409.07551
- Zhao X, Gao B. AI and microfluidics: Unlocking cellular motility for bioengineering. Bioengineering. 2026;13(2):172. doi: 10.3390/bioengineering13020172
- Cheng J, Anne R, Chen YC. Transforming microfluidics for single-cell analysis with robotics and artificial intelligence. Lab Chip. 2025;25(23):6100-6125. doi: 10.1039/d5lc00216h
- Gao Z, Li Y. Enhancing single-cell biology through advanced AI-powered microfluidics. Biomicrofluidics. 2023;17(5):051301. doi: 10.1063/5.0170050
- Mojica-Perez SP, Stokes K, Jaklic DC, et al. Protocol for selecting single human pluripotent stem cells using a modified micropipetter. STAR Protoc. 2023;4(4):102629. doi: 10.1016/j.xpro.2023.102629
- Garg G, Patel P, Gupta GD, Kurmi BD. A Review on Working Principle and Advanced Applications of Fluorescence activated Cell Sorting Machine (FACS). Curr Pharm Anal. 2024;20(2):85-97. doi: 10.2174/0115734129301463240306064929
- Liu D, Sun M, Zhang J, et al. Single-cell droplet microfluidics for biomedical applications. Analyst. 2022;147(11):2294-2316. doi: 10.1039/D1AN02321G
- Li B, Ma X, Cheng J, et al. Droplets microfluidics platform—A tool for single cell research. Front Bioeng Biotechnol. 2023;11:1121870. doi: 10.3389/fbioe.2023.1121870
- Ali M, Kim W, Park J. Droplet microfluidic technologies for next-generation high throughput screening. Front Lab Chip Technol. 2023;2:1230791. doi: 10.3389/frlct.2023.1230791
- Yuan J, Sims PA. An automated microwell platform for large-scale single cell RNA-seq. Sci Rep. 2016;6(1):33883. doi: 10.1038/srep33883
- Gebreyesus ST, Siyal AA, Kitata RB, et al. Streamlined single-cell proteomics by an integrated microfluidic chip and data-independent acquisition mass spectrometry. Nat Commun. 2022;13(1):37. doi: 10.1038/s41467-021-27778-4
- Fan X, Yang C, Li W, et al. SMOOTH-seq: single-cell genome sequencing of human cells on a third-generation sequencing platform. Genome Biol. 2021;22(1):195. doi: 10.1186/s13059-021-02406-y
- Tang J, Tu K, Lu K, et al. Single-cell exome sequencing reveals multiple subclones in metastatic colorectal carcinoma. Genome Med. 2021;13(1):148. doi: 10.1186/s13073-021-00962-3
- Hernandez-Hernandez A, Hojer P, Ljungmark M, et al. Comparative analysis of whole genome amplification kits for single cell genome analysis. bioRxiv. Preprint posted online October 31, 2025. doi: 10.1101/2025.10.30.685509
- Gonzalez-Pena V, Natarajan S, Xia Y, et al. Accurate genomic variant detection in single cells with primary template-directed amplification. Proc Natl Acad Sci USA. 2021;118(24):e2024176118. doi: 10.1073/pnas.2024176118
- Shi P, Nie Y, Yang J, Zhang W, Tang Z, Xu J. Fundamental and practical approaches for single-cell ATAC-seq analysis. Abiotech. 2022;3(3):212-223. doi: 10.1007/s42994-022-00082-5
- Shen Q, Deng E, Luo L, et al. High‐throughput single‐cell DNA methylation and chromatin accessibility co‐profiling with SpliCOOL‐seq. Clin Transl Med. 2026;16(2). doi: 10.1002/ctm2.70584
- Li G, Liu Y, Zhang Y, et al. Joint profiling of DNA methylation and chromatin architecture in single cells. Nat Methods. 2019;16(10):991-993. doi: 10.1038/s41592-019-0502-z
- De Rop FV, Hulselmans G, Flerin C, et al. Systematic benchmarking of single-cell ATAC-sequencing protocols. Nat Biotechnol. 2024;42(6):916-926. doi: 10.1038/s41587-023-01881-x
- Kashima Y, Sakamoto Y, Kaneko K, Seki M, Suzuki Y, Suzuki A. Single-cell sequencing techniques from individual to multiomics analyses. Exp Mol Med. 2020;52(9):1419-1427. doi: 10.1038/s12276-020-00499-2
- Stoeckius M, Hafemeister C, Stephenson W, et al. Simultaneous epitope and transcriptome measurement in single cells. Nat Methods. 2017;14(9):865-868. doi: 10.1038/nmeth.4380
- Williams CG, Lee HJ, Asatsuma T, Vento-Tormo R, Haque A. An introduction to spatial transcriptomics for biomedical research. Genome Med. 2022;14(1):68. doi: 10.1186/s13073-022-01075-1
- Moses L, Pachter L. Museum of spatial transcriptomics. Nat Methods. 2022;19(5):534-546. doi: 10.1038/s41592-022-01409-2
- Petrova B, Guler AT. Recent developments in single-cell metabolomics by mass Spectrometry: A perspective. J Proteome Res. 2024;24(4):1493-1518. doi: 10.1021/acs.jproteome.4c00646
- Zhang C, Le Devedec SE, Ali A, Hankemeier T. Single-cell metabolomics by mass spectrometry: Ready for primetime? Curr Opin Biotechnol. 2023;82:102963. doi: 10.1016/j.copbio.2023.102963
- Ma S, Zhang B, LaFave LM, et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell. 2020;183(4):1103-1116. doi: 10.1016/j.cell.2020.09.056
- Clark SJ, Argelaguet R, Kapourani CA, et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat Commun. 2018;9(1):781. doi: 10.1038/s41467-018-03149-4
- Lee J, Hyeon DY, Hwang D. Single-cell multiomics: Technologies and data analysis methods. Exp Mol Med. 2020;52(9):1428-1442. doi: 10.1038/s12276-020-0420-2
- Wu B, Bennett HM, Ye X, et al. Overloading And unpacking (OAK)-droplet-based combinatorial indexing for ultra-high throughput single-cell multiomic profiling. Nat Commun. 2024;15(1):9146. doi: 10.1038/s41467-024-53227-z
- Chow A, Lareau CA. Concepts and new developments in droplet-based single cell multi-omics. Trends Biotechnol. 2024;42(11):1379-1395. doi: 10.1016/j.tibtech.2024.07.006
- Brookes DH, Park H, Listgarten J. Conditioning by adaptive sampling for robust design. arXiv. Preprint posted online 2019. doi: 10.48550/arXiv.1901.10060
- Jeong Y, Ronen J, Kopp W, Lutsik P, Akalin A. scMaui: A widely applicable deep learning framework for single-cell multiomics integration in the presence of batch effects and missing data. BMC Bioinform. 2024;25:257. doi: 10.1186/s12859-024-05880-w
- Niu J, Vasquez-Rios C, Ding J. Single-cell multiomics data integration and generation with scPairing. bioRxiv. Preprint posted online January 4, 2025. doi: 10.1101/2025.01.04.631299
- Cheng Z, Lu R, Zhang S. scMGCL: Accurate and efficient integration representation of single-cell multi-omics data. Bioinformatics. 2025;41(7):btaf392. doi: 10.1093/bioinformatics/btaf392
- Argelaguet R, Arnol D, Bredikhin D, et al. MOFA+: A statistical framework for comprehensive integration of multi-modal single-cell data. Genome Biol. 2020;21(1):111. doi: 10.1186/s13059-020-02015-1
- Lopez R, Regier J, Cole MB, Jordan MI, Yosef N. Deep generative modeling for single-cell transcriptomics. Nat Methods. 2018;15(12):1053-1058. doi: 10.1038/s41592-018-0229-2.
- Ding J, Regev A. Deep generative model embedding of single-cell RNA-Seq profiles on hyperspheres and hyperbolic spaces. Nat Commun. 2021;12(1):2554. doi: 10.1038/s41467-021-22851-4.
- Perakis A, Gorji A, Jain S, Chaitanya K, Rizza S, Konukoglu E. Contrastive Learning of Single-Cell Phenotypic Representations for Treatment Classification. In: Machine Learning in Medical Imaging (Lecture Notes in Computer Science). Cham, Switzerland: Springer International Publishing; 2021:565-575. doi: 10.1007/978-3-030-87589-3_58
- Killoran N, Lee LJ, Delong A, Duvenaud D, Frey BJ. Generating and designing DNA with deep generative models. arXiv. Preprint posted online 2017. doi: 10.48550/arXiv.1712.06148
- Hao Y, Hao S, Andersen-Nissen E, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184(13):3573-3587. doi: 10.1016/j.cell.2021.04.048.
- Li T, Wang Z, Liu Y, He S, Zou Q, Zhang Y. An overview of computational methods in single-cell transcriptomic cell type annotation. Brief Bioinform. 2025;26(3). doi: 10.1093/bib/bbaf207
- Cui H, Wang C, Maan H, et al. scGPT: Toward building a foundation model for single-cell multi-omics using generative AI. Nat Methods. 2024;21(8):1470-1480. doi: 10.1038/s41592-024-02201-0
- Hutton A, Meyer JG. Trajectory inference for single cell omics. arXiv. Preprint posted online 2025. doi: 10.48550/arXiv.2502.09354
- Chang Z, Xu Y, Dong X, Gao Y, Wang C. Single-cell and spatial multiomic inference of gene regulatory networks using SCRIPro. Bioinformatics. 2024;40(7):btae466. doi: 10.1093/bioinformatics/btae466
- Barylli M, Saha J, Buffart TE, et al. Biological Multi-Layer and Single Cell Network-Based Multiomics Models—A Review. arXiv. Preprint posted online 2025. doi: 10.48550/arXiv.2503.09568
- Yang X, Mann KK, Wu H, Ding J. scCross: A deep generative model for unifying single-cell multi-omics with seamless integration, cross-modal generation, and in silico exploration. Genome Biol. 2024;25:198. doi: 10.1186/s13059-024-03338-z
- Li L, Dong L, Zhang H, Xu D, Li Y. spaLLM: Enhancing spatial domain analysis in multi-omics data through large language model integration. Brief Bioinform. 2025;26(4):bbaf304. doi: 10.1093/bib/bbaf304
- Nauwynck W, Faust K, Boon N. Droplet microfluidics for single-cell studies: a frontier in ecological understanding of microbiomes. FEMS Microbiol Rev. 2025;49:fuaf032. doi: 10.1093/femsre/fuaf032
- Sadasivan S, Pradeep S, Ramachandran JC, Narayan J, Gęca MJ. Advances in droplet microfluidics: A comprehensive review of innovations, morphology, dynamics, and applications. Microfluid Nanofluid. 2025;29(3):17. doi: 10.1007/s10404-025-02789-5
- Yi C, Cheng J, Chen J, Liu W, Liu J, Li Y. Benchmarking deep learning methods for biologically conserved single-cell integration. Genome Biol. 2025;26:398. doi: 10.1186/s13059-025-03869-z
- Yiu T, Chen B, Wang H, Feng G, Fu Q, Hu H. Transformative advances in single-cell omics: a comprehensive review of foundation models, multimodal integration and computational ecosystems. J Transl Med. 2025;23(1):1176. doi: 10.1186/s12967-025-07091-0
- Lambert CLG, van Mierlo G, Bues JJ, Guillaume-Gentil OJ, Deplancke B. Engineering next-generation microfluidic technologies for single-cell phenomics. Nat Genet. 2025;57(6):1344-1356. doi: 10.1038/s41588-025-02198-y
- Middlebrooks SA, Zhao X, Ford RM, Cummings PT. A mathematical model for Escherichia coli chemotaxis to competing stimuli. Biotechnol Bioeng. 2021;118(12):4678-4686. doi: 10.1002/bit.27930
- Ashuach T, Gabitto MI, Koodli RV, et al. MultiVI: Deep generative model for the integration of multimodal data. Nat Methods. 2023;20(8):1222-1231. doi: 10.1038/s41592-023-01909-9
- Song HW, Martin J, Shi X, Tyznik AJ. Key Considerations on CITE‐Seq for Single‐Cell Multiomics. Proteomics. 2025:e202400011. doi: 10.1002/pmic.202400011
- Sarfraz I, Wang Y, Shastry A, et al. MAMS: matrix and analysis metadata standards to facilitate harmonization and reproducibility of single-cell data. Genome Biol. 2024;25(1):205. doi: 10.1186/s13059-024-03349-w
- Yuan R, Rong Z, Hu H, et al. Harmony-based data integration for distributed single-cell multi-omics data. PLoS Comput Biol. 2025;21(9):e1013526. doi: 10.1371/journal.pcbi.1013526
- Fu S, Wang S, Si D, et al. Benchmarking single-cell multimodal data integrations. Nat Methods. 2025;22:2437–2448. doi: 10.1038/s41592-025-02737-9
- Wagle MM, Long S, Chen C, et al. Interpretable deep learning in singlecell omics. Bioinformatics. 2024;40(6):btae374. doi: 10.1093/bioinformatics/btae374
- Fessenko DS, Jasperse A. Ethics at the heart of AI regulation. AI Ethics. 2025;5:3387–3398. doi: 10.1007/s43681-024-00562-y
- Kumari P, Kaur M, Dindhoria K, et al. Advances in longread singlecell transcriptomics. Hum Genet. 2024;143(9-10):1005–1020. doi: 10.1007/s00439-024-02678-x
- Richards M, Malic N, Osterlund EJ, et al. Multiplexed confocal FLIM for dynamic molecular imaging in live cells. Discover Imaging. 2025;2(1):7. doi: 10.1007/s44352-025-00010-5

