M2Echem: A multilevel dual encoder-based model for predicting organic chemistry reactions

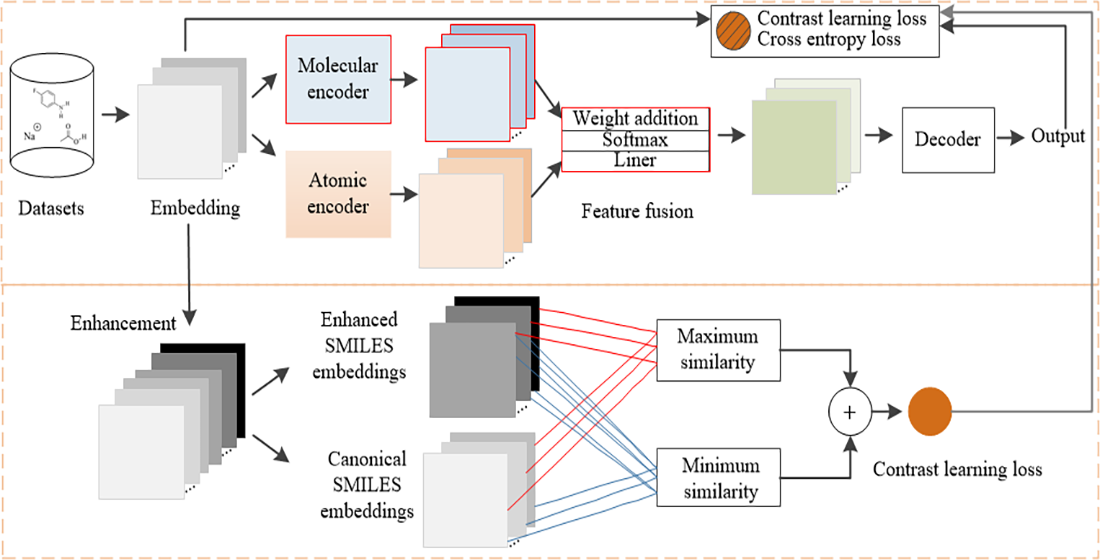
Chemical reaction prediction is a vital application of artificial intelligence. While Transformer models are widely used for this task, they often overlook deeper-level semantic information. In addition, the traditional Transformer model suffers from a decline in prediction performance and shows poor generalization when faced with different representations of the same molecule. To address these challenges, we propose a dual encoder-based reaction prediction method tailored for multilevel organic chemistry. Our approach began with the introduction of synergistic dual-encoder architecture: The atomic encoder focused on inter-atomic attention weights. In contrast, the molecular encoder employed a molecular maximum dimension reduction algorithm to identify key chemical features. We then performed multilevel feature fusion by combining the outputs from both the atomic and molecular encoders. Finally, we applied an optimized contrast loss to enhance the model’s robustness. The results indicated that this method outperformed existing models across all four datasets, significantly improving generalization performance and contributing to advancements in artificial intelligence-driven drug development and research.
- Corey EJ, Wipke WT. Computer-assisted design of complex organic syntheses. Science. 1969;166(3902):178-192. doi: 10.1126/science.166.3902.178
- Satoh H, Funatsu K. Further development of a reaction generator in the SOPHIA system for organic reaction prediction. Knowledge-guided addition of suitable atoms and/or atomic groups to product skeleton. J Chem Inform Comput Sci. 1996;36(2):173-184. doi: 10.1021/ci950058a
- Coley CW, Barzilay R, Jaakkola TS, Green WH, Jensen KF. Prediction of organic reaction outcomes using machine learning. ACS Cent Sci. 2017;3(5):434-443. doi: 10.1021/acscentsci.7b00064
- Duvenaud DK, Maclaurin D, Iparraguirre J, et al. Convolutional networks on graphs for learning molecular fingerprints. In: Proceedings of the 29th International Conference on Neural Information Processing Systems. Volume 2. California: Curran Associates, Inc.; 2015:2224-2232.
- Raccuglia P, Elbert KC, Adler PD, et al. Machine-learning-assisted materials discovery using failed experiments. Nature. 2016;533(7601):73-76. doi: 10.1038/nature17439
- Segler MH, Waller MP. Modelling chemical reasoning to predict and invent reactions. Chemistry. 2017;23(25):6118-6128. doi: 10.1002/chem.201604556
- Weininger DJ. SMILES, a chemical language and information system. 1. Introduction to methodology and encoding rules. J Chem Infrom Comput Sci. 1988;28(1):31-36. doi: 10.1021/ci00057a005
- Schwaller P, Gaudin T, Lanyi D, Bekas C, Laino TJ. “Found in translation”: Predicting outcomes of complex organic chemistry reactions using neural sequence-to-sequence models. Chem Sci. 2018;9(28):6091-6098. doi: 10.1039/c8sc02339e
- Vaswani A, Shazeer N, Parmar N, et al. Attention is All you Need. In: Proceedings of the 31st International Conference on Neural Information Processing Systems. 2017:6000-6010.
- Schwaller P, Laino T, Gaudin T, et al. Molecular transformer: A model for uncertainty-calibrated chemical reaction prediction. ACS Cent Sci. 2019;5(9):1572-1583. doi: 10.1021/acscentsci.9b00576
- Tang G, Müller M, Rios A, Sennrich R. Why Self-Attention? A Targeted Evaluation of Neural Machine Translation Architectures. In: Proceedings of the 2018 Conference on Empirical Methods in Natural Language Processing. Association for Computational Linguistics; 2018:4263-4272. doi: 10.18653/v1/d18-1458
- Wu F, Fan A, Baevski A, Dauphin YN, Auli M. Pay Less Attention with Lightweight and Dynamic Convolutions. arXiv. Preprint posted online 2019. doi: 10.48550/arXiv.1901.10430
- Schwaller P, Probst D, Vaucher AC, et al. Mapping the space of chemical reactions using attention-based neural networks. Nat Mach Intell. 2021;3(2):144-152. doi: 10.1038/s42256-020-00284-w
- Mellah Y, Kocaman V, Haq HU, Talby D. Efficient schema-less text-to-SQL conversion using large language models. Artif Intell Health. 2024;1(2):96-106. doi: 10.36922/aih.2661
- Mumtaz U, Ahmed A, Mumtaz S. LLMs-Healthcare: Current applications and challenges of large language models in various medical specialties. Artif Intell Health. 2024;1(2):16-28. doi: 10.36922/aih.2558
- Bran AM, Schwaller P. Transformers and large language models for chemistry and drug discovery. In: Drug Development Supported by Informatics. Berlin: Springer; 2024. p. 143-163.
- Leon M, Perezhohin Y, Peres F, Popovič A, Castelli M. Comparing SMILES and SELFIES tokenization for enhanced chemical language modeling. Sci Rep. 2024;14(1):25016. doi: 10.1038/s41598-024-76440-8
- Xiong J, Zhang W, Wang Y, et al. Bridging chemistry and artificial intelligence by a reaction description language. Nat Mach Intell. 2025;7(5):782-793. doi: 10.1038/s42256-025-01032-8
- Lo A, Pollice R, Nigam A, White AD, Krenn M, Aspuru- Guzik AJ. Recent advances in the self-referencing embedded strings (SELFIES) library. Dig Discov. 2023;2(4):897-908. doi: 10.1039/D3DD00044C
- Ucak UV, Ashyrmamatov I, Lee J. Improving the quality of chemical language model outcomes with atom-in-SMILES tokenization. J Cheminform. 2023;15(1):55. doi: 10.1186/s13321-023-00725-9
- Wu Z, Jiang D, Wang J, et al. Knowledge-based BERT: A method to extract molecular features like computational chemists. Brief Bioinform. 2022;23(3):bbac131. doi: 10.1093/bib/bbac131
- Chen S, Jung Y. Deep retrosynthetic reaction prediction using local reactivity and global attention. JACS Au. 2021;1(10):1612-1620. doi: 10.1021/jacsau.1c00246
- Liu Z, Zhang W, Xia Y, et al. MolXPT: Wrapping Molecules with Text for Generative Pre-training. In: Proceedings of the 61st Annual Meeting of the Association for Computational Linguistics (Volume 2: Short Papers). Association for Computational Linguistics; 2023:1606-1616. doi: 10.18653/v1/2023.acl-short.138
- Lu J, Zhang Y. Unified deep learning model for multitask reaction predictions with explanation. J Chem Inform Model. 2022;62(6):1376-1387. doi: 10.1021/acs.jcim.1c01467
- Guo W, Wang J, Wang S. Deep multimodal representation learning: A survey. IEEE Access. 2019;7:63373-63394. doi: 10.1109/ACCESS.2019.2916887
- Ma S, Zhang D, Zhou M. A Simple and Effective Unified Encoder for Document-Level Machine Translation. In: Proceedings of the 58th Annual Meeting of the Association for Computational Linguistics. Association for Computational Linguistics; 2020:3505-3511. doi: 10.18653/v1/2020.acl-main.321
- Zhang X, Li P, Li H. AMBERT: A Pre-trained Language Model with Multi-Grained Tokenization. arXiv. Preprint posted online 2020. doi: 10.48550/arXiv.2008.11869
- Zhu J, Xia Y, Wu L, et al. Incorporating BERT into Neural Machine Translation. arXiv. Preprint posted online 2020. doi: 10.48550/arXiv.2002.06823
- Jin D, Jin Z, Zhou JT, Szolovits P. Is BERT Really Robust? A Strong Baseline for Natural Language Attack on Text Classification and Entailment. Proceedings of the AAAI Conference on Artificial Intelligence. 2020;34(05):8018-8025. doi: 10.1609/aaai.v34i05.6311
- Singh S, Shingatgeri V, Srivastava P. Revolutionizing new drug discovery: Harnessing AI and machine learning to overcome traditional challenges and accelerate targeted therapies. Artif Intell Health. 2024;2(2):29-40. doi: 10.36922/aih.4423
- Gao T, Yao X, Chen D. SimCSE: Simple Contrastive Learning of Sentence Embeddings. arXiv. Preprint posted online 2021. doi: 10.48550/arXiv.2104.08821
- Chen X, Alamro H, Li M, et al. Target-aware Abstractive Related Work Generation with Contrastive Learning. In: Proceedings of the 45th International ACM SIGIR Conference on Research and Development in Information Retrieval. Association for Computing Machinery; 2022:373-383. doi: 10.1145/3477495.3532065
- Miculicich L, Ram D, Pappas N, Henderson J. Document- Level Neural Machine Translation with Hierarchical Attention Networks. In: Proceedings of the 2018 Conference on Empirical Methods in Natural Language Processing. Belgium: Association for Computational Linguistics; 2018:2947–2954.
- Mao A, Mohri M, Zhong Y. Cross-Entropy Loss Functions: Theoretical Analysis and Applications. In: Proceedings of the 40th International Conference on Machine Learning. 2023:23803-23828.
- Jiang S, Zhang Z, Zhao H, et al. When SMILES smiles, practicality judgment and yield prediction of chemical reaction via deep chemical language processing. IEEE Access. 2021;9:85071-85083. doi: 10.1109/ACCESS.2021.3083838
- Lowe D. Chemical reactions from US patents (1976- Sep2016). Published online 2017. doi: 10.6084/M9.FIGSHARE.5104873
- Liu B, Ramsundar B, Kawthekar P, et al. Retrosynthetic reaction prediction using neural sequence-to-sequence models. ACS Cent Sci. 2017;3(10):1103-1113. doi: 10.1021/acscentsci.7b00303
- Jin W, Coley C, Barzilay R, Jaakkola TJ. Predicting Organic Reaction Outcomes with Weisfeiler-Lehman Network. In: Proceedings of the 31st International Conference on Neural Information Processing Systems. 2017:2604-2613.
- Papineni K, Roukos S, Ward T, Zhu WJ. BLEU. In: Proceedings of the 40th Annual Meeting on Association for Computational Linguistics - ACL ’02. Association for Computational Linguistics; 2001:311-318. doi: 10.3115/1073083.1073135
- Wang T. Research on chemical reaction prediction model based on Fairseq. In: 2021 2nd International Conference on Big Data & Artificial Intelligence & Software Engineering (ICBASE). IEEE; 2021:167-171. doi: 10.1109/icbase53849.2021.00039
- Bjerrum EJ. SMILES Enumeration as Data Augmentation for Neural Network Modeling of Molecules. arXiv. Preprint posted online 2017. doi: 10.48550/arXiv.1703.07076
- Tetko IV, Karpov P, Van Deursen R, Godin G. State-of-the-art augmented NLP transformer models for direct and single-step retrosynthesis. Nat Commun. 2020;11(1):5575. doi: 10.1038/s41467-020-19266-y
- Khalifa AA, Haranczyk M, Holliday J. Comparison of nonbinary similarity coefficients for similarity searching, clustering and compound selection. J Chem Inf Model. 2009;49(5):1193-1201. doi: 10.1021/ci8004644

