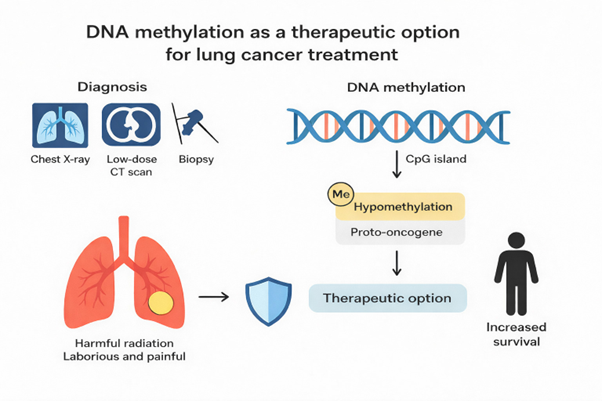
The function of DNA methylation in lung cancer: Diagnostic and clinical implications

Currently, the detection of lung cancer depends on chest X-ray, low-dose computed tomography (CT) scans, and tissue biopsy. However, harmful radiation from X-rays and CT scans may increase cancer risk. In contrast, biopsy of the lungs is highly laborious and painful. These diagnostic limitations contribute to delayed diagnosis, and treatment is therefore less effective at advanced stages. DNA methylation is a mechanism in which DNA methyltransferases add a methyl group to the cytosine–phosphate–guanine islands of a gene, which can either enhance or suppress its expression. Previous research has shown that promoter methylation of protooncogenes and tumor suppressor genes begins at the early stages of tumorigenesis and can be detected in cell-free DNA derived from blood plasma. With this primary objective, we highlight DNA hypomethylation in proto-oncogenes and hypermethylation in tumor suppressor genes in lung cancer patients, which may serve as therapeutic targets for the development of treatments that may increase the survival rate of lung cancer patients.
- Rudin CM, Brambilla E, Faivre-Finn C, Sage J. Small-cell lung cancer. Nat Rev Dis Primers. 2021;7(1):3. doi: 10.1038/s41572-020-00235-0
- Schübeler D. Function and information content of DNA methylation. Nature. 2015;517(7534):321-326. doi: 10.1038/nature14192
- Karamitrousis EI, Balgkouranidou I, Xenidis N, et al. Prognostic Role of RASSF1A, SOX17 and Wif-1 Promoter Methylation Status in Cell-Free DNA of Advanced Gastric Cancer Patients. Technol Cancer Res Treat. 2021;20. doi: 10.1177/1533033820973279
- Chen E, Cario CL, Leong L, et al. Cell-Free DNA Detection of Tumor Mutations in Heterogeneous, Localized Prostate Cancer Via Targeted, Multiregion Sequencing. JCO Precis Oncol. 2021;(5):710-725. doi: 10.1200/PO.20.00428
- Mouliere F, Chandrananda D, Piskorz AM, et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci Transl Med. 2018;10(466). doi: 10.1126/scitranslmed.aat4921
- Zhao Y, Yang M, Wang S, et al. An Overview of Epigenetic Methylation in Pancreatic Cancer Progression. Front Oncol. 2022;12:854773. doi: 10.3389/fonc.2022.854773
- Du Q, Luu PL, Stirzaker C, Clark SJ. Methyl-CpG-binding domain proteins: readers of the epigenome. Epigenomics. 2015;7(6):1051-1073. doi: 10.2217/epi.15.39
- Vaughan RM, Dickson BM, Cornett EM, Harrison JS, Kuhlman B, Rothbart SB. Comparative biochemical analysis of UHRF proteins reveals molecular mechanisms that uncouple UHRF2 from DNA methylation maintenance. Nucleic Acids Res. 2018;46(9):4405-4416. doi: 10.1093/nar/gky151
- Abu-Alainin W, Gana T, Liloglou T, et al. UHRF1 regulation of the Keap1-Nrf2 pathway in pancreatic cancer contributes to oncogenesis. J Pathol. 2016;238(3):423-433. doi: 10.1002/path.4665
- Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP. Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res. 1998;72:141-196. doi: 10.1016/S0065-230X(08)60702-2
- Kulis M, Esteller M. DNA methylation and cancer. Adv Genet. 2010;70:27-56. doi: 10.1016/B978-0-12-380866-0.60002-2
- Chen T, Li E. Establishment and maintenance of DNA methylation patterns in mammals. Curr Top Microbiol Immunol. 2006;301:179-201. doi: 10.1007/3-540-31390-7_6
- Belinsky SA, Klinge DM, Stidley CA, et al. Inhibition of DNA methylation and histone deacetylation prevents murine lung cancer. Cancer Res. 2003;63(21):7089-7093.
- Kassis ES, Zhao M, Hong JA, Chen GA, Nguyen DM, Schrump DS. Depletion of DNA methyltransferase 1 and/or DNA methyltransferase 3b mediates growth arrest and apoptosis in lung and esophageal cancer and malignant pleural mesothelioma cells. J Thorac Cardiovasc Surg. 2006;131(2):298-306. doi: 10.1016/j.jtcvs.2005.05.022
- Fabbri M, Garzon R, Cimmino A, et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc Natl Acad Sci USA. 2007;104(40):15805-15810. doi: 10.1073/pnas.0707628104
- Husni RE, Shiba-Ishii A, Iiyama S, et al. DNMT3a expression pattern and its prognostic value in lungadenocarcinoma. Lung Cancer. 2016;97:59-65. doi: 10.1016/j.lungcan.2016.04.018
- Sato M, Horio Y, Sekido Y, Minna JD, Shimokata K, Hasegawa Y. The expression of DNA methyltransferases and methyl-CpG-binding proteins is not associated with the methylation status of p14(ARF), p16(INK4a) and RASSF1A in human lung cancer cell lines. Oncogene. 2002;21(31):4822-4829. doi: 10.1038/sj.onc.1205581
- Wu XY, Chen HC, Li WW, Yan JD, Lv RY. DNMT1 promotes cell proliferation via methylating hMLH1 and hMSH2 promoters in EGFR-mutated non-small cell lung cancer. J Biochem. 2020;168(2):151-157. doi: 10.1093/jb/mvaa034
- Mutskov VJ, Farrell CM, Wade PA, Wolffe AP, Felsenfeld G. The barrier function of an insulator couples high histone acetylation levels with specific protection of promoter DNA from methylation. Genes Dev. 2002;16(12):1540-1554. doi: 10.1101/gad.988502
- Frigola J, Song J, Stirzaker C, Hinshelwood RA, Peinado MA, Clark SJ. Epigenetic remodeling in colorectal cancer results in coordinate gene suppression across an entire chromosome band. Nat Genet. 2006;38(5):540-549. doi: 10.1038/ng1781
- Serrano M, Hannon GJ, Beach D. A new regulatory motif in cell-cycle control causing specific inhibition of cyclin D/CDK4. Nature. 1993;366(6456):704-707. doi: 10.1038/366704a0
- Singhal S, Vachani A, Antin-Ozerkis D, Kaiser LR, Albelda SM. Prognostic implications of cell cycle, apoptosis, and angiogenesis biomarkers in non-small cell lung cancer: a review. Clin Cancer Res. 2005;11(11):3974-3986. doi: 10.1158/1078-0432.CCR-04-2661
- Aravidis C, Panani AD, Kosmaidou Z, Thomakos N, Rodolakis A, Antsaklis A. Detection of numerical abnormalities of chromosome 9 and p16/CDKN2A gene alterations in ovarian cancer with fish analysis. Anticancer Res. 2012;32(12):5309-5313.
- Cairns P, Polascik TJ, Eby Y, et al. Frequency of homozygous deletion at p16/CDKN2 in primary human tumours. Nat Genet. 1995;11(2):210-212. doi: 10.1038/ng1095-210
- Liu Y, An Q, Li L, et al. Hypermethylation of p16INK4a in Chinese lung cancer patients: biological and clinical implications. Carcinogenesis. 2003;24(12):1897-1901. doi: 10.1093/carcin/bgg169
- Ostrow KL, Hoque MO, Loyo M, et al. Molecular analysis of plasma DNA for the early detection of lung cancer by quantitative methylation-specific PCR. Clin Cancer Res. 2010;16(13):3463-3472. doi: 10.1158/1078-0432.CCR-09-3304
- Balgkouranidou I, Chimonidou M, Milaki G, et al. Breast cancer metastasis suppressor-1 promoter methylation in cell-free DNA provides prognostic information in nonsmall cell lung cancer. Br J Cancer. 2014;110(8):2054-2062. doi: 10.1038/bjc.2014.104
- Fu DY, Wang ZM, Li C, et al. Sox17, the canonical Wnt antagonist, is epigenetically inactivated by promoter methylation in human breast cancer. Breast Cancer Res Treat. 2010;119(3):601-612. doi: 10.1007/s10549-009-0339-8
- Ehrlich M. DNA hypomethylation in cancer cells. Epigenomics. 2009;1(2):239-259. doi: 10.2217/epi.09.33
- Vachtenheim J, Horáková I, Novotná H. Hypomethylation of CCGG sites in the 3’ region of H-ras protooncogene is frequent and is associated with H-ras allele loss in non-small cell lung cancer. Cancer Res. 1994;54(5):1145-1148.
- Yin J, Hou W, Vogel U, et al. TP53 common variants and interaction with PPP1R13L and CD3EAP SNPs and lung cancer risk and smoking behavior in a Chinese population. Biomed J. 2022;45(1):169-178. doi: 10.1016/j.bj.2021.01.006
- Belinsky SA, Grimes MJ, Casas E, et al. Predicting gene promoter methylation in non-small-cell lung cancer by evaluating sputum and serum. Br J Cancer. 2007;96(8):1278-1283. doi: 10.1038/sj.bjc.6603721
- Krishnamurthy K, Mishra TK, Saxena A, et al. Evaluating NISCH and CDH1 Promoter Hypermethylation in Nonsmokers, Cancer Free Smokers and Lung Cancer Patients: A Case Control Study. Indian J Clin Biochem. 2019;34(4):458-464. doi: 10.1007/s12291-018-0767-5
- Sadri F, Alipour M, Nejatizadeh A. Significant Association Among the Epigenetic Alterations in EGFR and ALK Genes and Non-Small Cell Lung Cancer: A Prospective Emerging Molecular Biomarker. Ann Hum Genet. 2025;89(6):425-433. doi: 10.1111/ahg.12597
- Gao Q, Steine EJ, Barrasa MI, et al. Deletion of the de novo DNA methyltransferase Dnmt3a promotes lung tumorprogression. Proc Natl Acad Sci USA. 2011;108(44):18061-18066. doi: 10.1073/pnas.1114946108
- Zhang C, Yu W, Wang L, et al. DNA Methylation Analysis of the SHOX2 and RASSF1A Panel in Bronchoalveolar LavageFluid for Lung Cancer Diagnosis. J Cancer. 2017;8(17):3585-3591. doi: 10.7150/jca.21368
- Baglietto L, Ponzi E, Haycock P, et al. DNA methylation changes measured in pre-diagnostic peripheral blood samples are associated with smoking and lung cancer risk. Int J Cancer. 2017;140(1):50-61. doi: 10.1002/ijc.30431
- Pulling LC, Divine KK, Klinge DM, et al. Promoter hypermethylation of the O6-methylguanine-DNA methyltransferase gene: more common in lung adenocarcinomas from never-smokers than smokers and associated with tumor progression. Cancer Res. 2003;63(16):4842-4848.
- Liu J, Yang XY, Shi WJ. Identifying differentially expressed genes and pathways in two types of non-small cell lung cancer: adenocarcinoma and squamous cell carcinoma. Genet Mol Res. 2014;13(1):95-102. doi: 10.4238/2014.January.8.8
- Pesek M, Kopeckova M, Benesova L, et al. Clinical significance of hypermethylation status in NSCLC: evaluation of a 30-gene panel in patients with advanced disease. Anticancer Res. 2011;31(12):4647-4652.
- Tsuboi M, Kondo K, Soejima S, et al. Chromate exposure induces DNA hypermethylation of the mismatch repair gene MLH1 in lung cancer. Mol Carcinog. 2020;59(1):24-31. doi: 10.1002/mc.23125
- Ali AH, Kondo K, Namura T, et al. Aberrant DNA methylation of some tumor suppressor genes in lung cancers from workers with chromate exposure. Mol Carcinog. 2011;50(2):89-99. doi: 10.1002/mc.20697
- Hu G, Li P, Cui X, et al. Cr(VI)-induced methylation and down-regulation of DNA repair genes and its association with markers of genetic damage in workers and 16HBE cells. Environ Pollut. 2018;238:833-843. doi: 10.1016/j.envpol.2018.03.046
- Hung CS, Wang SC, Yen YT, Lee TH, Wen WC, Lin RK. Hypermethylation of CCND2 in Lung and Breast Cancer Is a Potential Biomarker and Drug Target. Int J Mol Sci. 2018;19(10):3096. doi: 10.3390/ijms19103096
- Castro M, Grau L, Puerta P, et al. Multiplexed methylation profiles of tumor suppressor genes and clinical outcome in lung cancer. J Transl Med. 2010;8(1). doi: 10.1186/1479-5876-8-86
- Oshimo Y, Nakayama H, Ito R, et al. Promoter methylation of cyclin D2 gene in gastric carcinoma. Int J Oncol. 2003;23(6):1663-1670.
- Wang J, Zhao Y, Xu H, et al. Silencing NID2 by DNA Hypermethylation Promotes Lung Cancer. Pathol Oncol Res. 2020;26(2):801-811. doi: 10.1007/s12253-019-00609-0
- Geng J, Sun J, Lin Q, et al. Methylation status of NEUROG2 and NID2 improves the diagnosis of stage I NSCLC. Oncol Lett. 2012;3(4):901-906. doi: 10.3892/ol.2012.587
- Su C, Shi K, Cheng X, et al. Methylation of CLEC14A is associated with its expression and lung adenocarcinoma progression. J Cell Physiol. 2019;234(3):2954-2962. doi: 10.1002/jcp.27112
- Ki MK, Jeoung MH, Choi JR, et al. Human antibodies targeting the C-type lectin-like domain of the tumor endothelial cell marker clec14a regulate angiogenic properties in vitro. Oncogene. 2013;32(48):5449-5457. doi: 10.1038/onc.2013.156
- Ramalho-Carvalho J, Henrique R, Jerónimo C. Methylation-Specific PCR. Methods Mol Biol. 2018;1708:447-472. doi: 10.1007/978-1-4939-7481-8_23
- Šestáková Š, Šálek C, Remešová H. DNA Methylation Validation Methods: a Coherent Review with Practical Comparison. Biol Proced Online. 2019;21:19. doi: 10.1186/s12575-019-0107-z
- Martisova A, Holcakova J, Izadi N, Sebuyoya R, Hrstka R, Bartosik M. DNA Methylation in Solid Tumors: Functions and Methods of Detection. Int J Mol Sci. 2021;22(8):4247. doi: 10.3390/ijms22084247
- Kwon YJ, Lee SJ, Koh JS, et al. Genome-wide analysis of DNA methylation and the gene expression change in lung cancer. J Thorac Oncol. 2012;7(1):20-33. doi: 10.1097/JTO.0b013e3182307f62
- Laird PW, Jackson-Grusby L, Fazeli A, et al. Suppression of intestinal neoplasia by DNA hypomethylation. Cell. 1995;81(2):197-205. doi: 10.1016/0092-8674(95)90329-1
- Bender CM, Pao MM, Jones PA. Inhibition of DNA methylation by 5-aza-2’-deoxycytidine suppresses the growth of human tumor cell lines. Cancer Res. 1998;58(1):95-101.
- Huang MH, Chou YW, Li MH, et al. Epigenetic targeting DNMT1 of pancreatic ductal adenocarcinoma using interstitial control release biodegrading polymer reduced tumor growth through hedgehog pathway inhibition. Pharmacol Res. 2019;139:50-61. doi: 10.1016/j.phrs.2018.10.015
- Leone G, Voso MT, Teofili L, Lübbert M. Inhibitors of DNA methylation in the treatment of hematological malignancies and MDS. Clin Immunol. 2003;109(1):89-102. doi: 10.1016/s1521-6616(03)00207-9
- Stresemann C, Lyko F. Modes of action of the DNA methyltransferase inhibitors azacytidine and decitabine. Int J Cancer. 2008;123(1):8-13. doi: 10.1002/ijc.23607
- Li P, Liu S, Du L, Mohseni G, Zhang Y, Wang C. Liquid biopsies based on DNA methylation as biomarkers for the detection and prognosis of lung cancer. Clin Epigenetics. 2022;14(1):118. doi: 10.1186/s13148-022-01337-0
- Bremnes RM, Sirera R, Camps C. Circulating tumour-derived DNA and RNA markers in blood: a tool for early detection, diagnostics, and follow-up? Lung Cancer. 2005;49(1):1-12. doi: 10.1016/j.lungcan.2004.12.008
- van der Drift MA, Hol BE, Klaassen CH, et al. Circulating DNA is a non-invasive prognostic factor for survival in non-small cell lung cancer. Lung Cancer. 2010;68(2):283-287. doi: 10.1016/j.lungcan.2009.06.021
- Yang Z, Chen Q, Dong S, et al. Hypermethylated TAGMe as a universal-cancer-only methylation marker and its application in diagnosis and recurrence monitoring of urothelial carcinoma. J Transl Med. 2024;22(1):608. doi: 10.1186/s12967-024-05420-3
- Gaga M, Chorostowska-Wynimko J, Horváth I, et al. Validation of Lung EpiCheck, a novel methylation-based blood assay, for the detection of lung cancer in European and Chinese high-risk individuals. Eur Respir J. 2021;57(1):2002682. doi: 10.1183/13993003.02682-2020
- Begum S, Brait M, Dasgupta S, et al. An epigenetic marker panel for detection of lung cancer using cell-free serum DNA. Clin Cancer Res. 2011;17(13):4494-4503. doi: 10.1158/1078-0432.CCR-10-3436
- Kneip C, Schmidt B, Seegebarth A, et al. SHOX2 DNA methylation is a biomarker for the diagnosis of lung cancer in plasma. J Thorac Oncol. 2011;6(10):1632-1638. doi: 10.1097/JTO.0b013e318220ef9a
- Hu Y, Ulrich BC, Supplee J, et al. False-Positive Plasma Genotyping Due to Clonal Hematopoiesis. Clin Cancer Res. 2018;24(18):4437-4443. doi: 10.1158/1078-0432.CCR-18-0143
- Kenaan N, Hanna G, Sardini M, et al. Advances in early detection of non-small cell lung cancer: A comprehensive review. Cancer Med. 2024;13(18):e70156. doi: 10.1002/cam4.70156
- Li W, Liu JB, Hou LK, et al. Liquid biopsy in lung cancer: significance in diagnostics, prediction, and treatment monitoring. Mol Cancer. 2022;21(1). doi: 10.1186/s12943-022-01505-z
- Chang L, Li J, Zhang R. Liquid biopsy for early diagnosis of non-small cell lung carcinoma: recent research and detection technologies. Biochim Biophys Acta Rev Cancer. 2022;1877(3):188729. doi: 10.1016/j.bbcan.2022.188729

