Intestinal organoids: Reshaping the paradigm of inflammatory bowel disease research and therapy

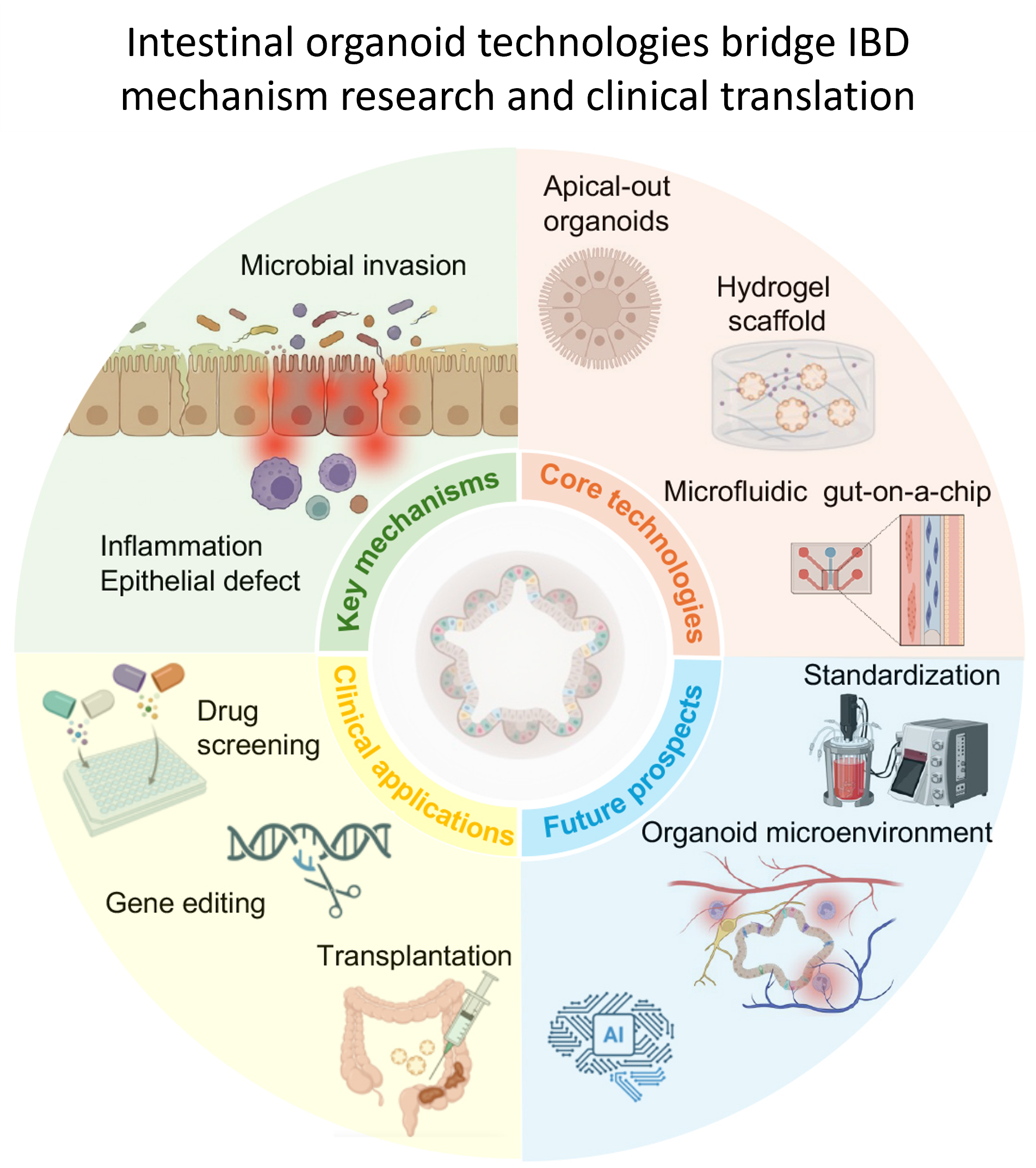
The advent of intestinal organoid technology has brought a revolutionary breakthrough to inflammatory bowel disease (IBD) research, enabling more precise in vitro simulation of human intestinal physiology and pathology. This article systematically reviews the latest advances in the application of intestinal organoid technology to IBD research. First, we introduce technological innovations such as synthetic hydrogels, microfluidic organ-on-a-chip systems, and polarity-reversal culture. These approaches overcome limitations of traditional two-dimensional cultures and greatly expand the applicability of organoid models. Second, we focus on research utilizing patient-derived organoids to unveil novel mechanisms of IBD pathogenesis, highlighting their unique advantages in dissecting intrinsic epithelial defects (e.g., impaired barrier function, abnormal mucus secretion) and dynamically simulating host–microbe interactions. Furthermore, we highlight clinical translation: high-throughput drug screening for personalized medicine and organoid transplantation for mucosal regeneration. Finally, we summarize current challenges regarding standardization, vascularization, and immune microenvironment reconstruction of organoids, and discuss their broad prospects for driving innovation in IBD diagnosis and treatment strategies.
- Ng SC, Shi HY, Hamidi N, et al. Worldwide incidence and prevalence of inflammatory bowel disease in the 21st century: a systematic review of population-based studies. Lancet. 2017;390(10114):2769-2778. doi: 10.1016/s0140-6736(17)32448-0
- Hirten RP, Iacucci M, Shah S, Ghosh S, Colombel JF. Combining Biologics in Inflammatory Bowel Disease and Other Immune Mediated Inflammatory Disorders. Clin Gastroenterol Hepatol. 2018;16(9):1374-1384. doi: 10.1016/j.cgh.2018.02.024
- Ahmed W, Galati J, Kumar A, et al. Dual Biologic or Small Molecule Therapy for Treatment of Inflammatory Bowel Disease: A Systematic Review and Meta-analysis. Clin Gastroenterol Hepatol. 2022;20(3):e361-e379. doi: 10.1016/j.cgh.2021.03.034
- Ramos GP, Papadakis KA. Mechanisms of Disease: Inflammatory Bowel Diseases. Mayo Clin Proc. 2019;94(1):155-165. doi: 10.1016/j.mayocp.2018.09.013
- Sazonovs A, Stevens CR, Venkataraman GR, et al. Large-scale sequencing identifies multiple genes and rare variants associated with Crohn’s disease susceptibility. Nat Genet. 2022;54(9):1275-1283. doi: 10.1038/s41588-022-01156-2
- VanDussen KL, Liu TC, Li D, et al. Genetic Variants Synthesize to Produce Paneth Cell Phenotypes That Define Subtypes of Crohn’s Disease. Gastroenterology. 2014;146(1):200-209. doi: 10.1053/j.gastro.2013.09.048
- Lin JD, Devlin JC, Yeung F, et al. Rewilding Nod2 and Atg16l1 Mutant Mice Uncovers Genetic and Environmental Contributions to Microbial Responses and Immune Cell Composition. Cell Host Microbe. 2020;27(5):830-840.e4. doi: 10.1016/j.chom.2020.03.001
- Kaya GG, Schwarzer R, Dannappel M, et al. Unfolded protein response transcription factor XBP1 suppresses necroptosis-induced colitis by reinforcing the mucus barrier. Immunity. 2025;58(9):2208-2225.e6. doi: 10.1016/j.immuni.2025.07.023
- Stengel ST, Fazio A, Lipinski S, et al. Activating Transcription Factor 6 Mediates Inflammatory Signals in Intestinal Epithelial Cells Upon Endoplasmic Reticulum Stress. Gastroenterology. 2020;159(4):1357-1374.e10. doi: 10.1053/j.gastro.2020.06.088
- Pierce R, Jan NJ, Kumar P, Middleton J, Petri WA, Marie C. Persistent dysbiosis of duodenal microbiota in patients with controlled pediatric Crohn’s disease after resolution of inflammation. Sci Rep. 2024;14(1):11567. doi: 10.1038/s41598-024-63299-y
- Rahmouni O, Vignal C, Titécat M, et al. High carriage of adherent invasive E. coli in wildlife and healthy individuals. Gut Pathog. 2018;10(1):23. doi: 10.1186/s13099-018-0248-7
- Pearce SC, Coia HG, Karl JP, Pantoja-Feliciano IG, Zachos NC, Racicot K. Intestinal in vitro and ex vivo Models to Study Host-Microbiome Interactions and Acute Stressors. Front Physiol. 2018;9:1584. doi: 10.3389/fphys.2018.01584
- Rahman S, Ghiboub M, Donkers JM, et al. The Progress of Intestinal Epithelial Models from Cell Lines to Gut-On- Chip. Int J Mol Sci. 2021;22(24):13472. doi: 10.3390/ijms222413472
- Sato T, Vries RG, Snippert HJ, et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature. 2009;459(7244):262-265. doi: 10.1038/nature07935
- Beumer J, Clevers H. Cell fate specification and differentiation in the adult mammalian intestine. Nat Rev Mol Cell Biol. 2020;22(1):39-53. doi: 10.1038/s41580-020-0278-0
- Nyström EEL, Martinez-Abad B, Arike L, et al. An intercrypt subpopulation of goblet cells is essential for colonic mucus barrier function. Science. 2021;372(6539):eabb1590. doi: 10.1126/science.abb1590
- Kollmann C, Buerkert H, Meir M, et al. Human organoids are superior to cell culture models for intestinal barrier research. Front Cell Dev Biol. 2023;11:1223032. doi: 10.3389/fcell.2023.1223032
- d’Aldebert E, Quaranta M, Sébert M, et al. Characterization of Human Colon Organoids From Inflammatory Bowel Disease Patients. Front Cell Dev Biol. 2020;8:363. doi: 10.3389/fcell.2020.00363
- Dennison TW, Edgar RD, Payne F, et al. Patient-derived organoid biobank identifies epigenetic dysregulation of intestinal epithelial MHC-I as a novel mechanism in severe Crohn’s Disease. Gut. 2024;73(9):1464-1477. doi: 10.1136/gutjnl-2024-332043
- Mao Y, Wang W, Yang J, et al. Drug repurposing screening and mechanism analysis based on human colorectal cancer organoids. Protein Cell. 2023;15(4):285-304. doi: 10.1093/procel/pwad038
- Hammerhøj A, Chakravarti D, Sato T, Jensen KB, Nielsen OH. Organoids as regenerative medicine for inflammatory bowel disease. iScience. 2024;27(6):110118. doi: 10.1016/j.isci.2024.110118
- Miura S, Suzuki A. Generation of Mouse and Human Organoid-Forming Intestinal Progenitor Cells by Direct Lineage Reprogramming. Cell Stem Cell. 2017;21(4):456- 471.e5. doi: 10.1016/j.stem.2017.08.020
- Sugimoto S, Ohta Y, Fujii M, et al. Reconstruction of the Human Colon Epithelium In Vivo. Cell Stem Cell. 2018;22(2):171-176.e5. doi: 10.1016/j.stem.2017.11.012
- Cadwell K, Loke P. Gene–environment interactions shape the host–microbial interface in inflammatory bowel disease. Nat Immunol. 2025;26(7):1023-1035. doi: 10.1038/s41590-025-02197-5
- Hirota A, AlMusawi S, Nateri AS, Ordóñez-Morán P, Imajo M. Biomaterials for intestinal organoid technology and personalized disease modeling. Acta Biomater. 2021;132:272- 287. doi: 10.1016/j.actbio.2021.05.010
- Kleinman HK, Martin GR. Matrigel: Basement membrane matrix with biological activity. Semin Cancer Biol. 2005;15(5):378-386. doi: 10.1016/j.semcancer.2005.05.004
- Cruz-Acuña R, Quirós M, Farkas AE, et al. Synthetic hydrogels for human intestinal organoid generation and colonic wound repair. Nat Cell Biol. 2017;19(11):1326-1335. doi: 10.1038/ncb3632
- Hughes CS, Postovit LM, Lajoie GA. Matrigel: A complex protein mixture required for optimal growth of cell culture. Proteomics. 2010;10(9):1886-1890. doi: 10.1002/pmic.200900758
- Capeling MM, Czerwinski M, Huang S, et al. Nonadhesive Alginate Hydrogels Support Growth of Pluripotent Stem Cell-Derived Intestinal Organoids. Stem Cell Reports. 2019;12(2):381-394. doi: 10.1016/j.stemcr.2018.12.001
- Gjorevski N, Sachs N, Manfrin A, et al. Designer matrices for intestinal stem cell and organoid culture. Nature. 2016;539(7630):560-564. doi: 10.1038/nature20168
- Vanhove L, Van Gansbeke T, Devriendt B, Van der Meeren R, Dmitriev RI, Okkelman IA. Lack of biochemical signalling in GelMA leads to polarity reversion in intestinal organoids independent from mechanoreciprocity. J Tissue Eng. 2025;16:20417314251345000. doi: 10.1177/20417314251345000
- Ma W, Zheng Y, Yang G, et al. A bioactive calcium silicate nanowire-containing hydrogel for organoid formation and functionalization. Mater Horiz. 2024;11(12):2957-2973. doi: 10.1039/d4mh00228h
- Madl CM, Heilshorn SC, Blau HM. Bioengineering strategies to accelerate stem cell therapeutics. Nature. 2018;557(7705):335-342. doi: 10.1038/s41586-018-0089-z
- Hushka EA, Blatchley MR, Macdougall LJ, et al. Fully Synthetic Hydrogels Promote Robust Crypt Formation in Intestinal Organoids. Adv Mater. 2025;37(43):e2509672. doi: 10.1002/adma.202509672
- He S, Lei P, Kang W, et al. Stiffness Restricts the Stemness of the Intestinal Stem Cells and Skews Their Differentiation Toward Goblet Cells. Gastroenterology. 2023;164(7):1137- 1151.e15. doi: 10.1053/j.gastro.2023.02.030
- Zhang Q, He J, Zhu D, et al. Genetically modified organoids for tissue engineering and regenerative medicine. Adv Colloid Interface Sci. 2025;335:103337. doi: 10.1016/j.cis.2024.103337
- Yang H, Feng R, Fu Q, et al. Human induced pluripotent stem cell-derived mesenchymal stem cells promote healing via TNF-α-stimulated gene-6 in inflammatory bowel disease models. Cell Death Dis. 2019;10(10):718. doi: 10.1038/s41419-019-1957-7
- Watanabe S, Nishimura R, Shirasaki T, et al. Schlafen 11 Is a Novel Target for Mucosal Regeneration in Ulcerative Colitis. J Crohns Colitis. 2021;15(9):1558-1572. doi: 10.1093/ecco-jcc/jjab032
- Xiao L, Ma XX, Luo J, et al. Circular RNA CircHIPK3 Promotes Homeostasis of the Intestinal Epithelium by Reducing MicroRNA 29b Function. Gastroenterology. 2021;161(4):1303-1317.e3. doi: 10.1053/j.gastro.2021.05.060
- D’Addio F, Amabile G, Assi E, et al. TMEM219 signaling promotes intestinal stem cell death and exacerbates colitis. J Clin Invest. 2025;135(10):e185783. doi: 10.1172/jci185783
- Kang Y, Park H, Choe BH, Kang B. The Role and Function of Mucins and Its Relationship to Inflammatory Bowel Disease. Front Med. 2022;9:848344. doi: 10.3389/fmed.2022.848344
- Ding W, Marx OM, Mankarious MM, Koltun WA, Yochum GS. Disease Severity Impairs Generation of Intestinal Organoid Cultures From Inflammatory Bowel Disease Patients. J Surg Res. 2024;293:187-195. doi: 10.1016/j.jss.2023.08.027
- Koslowski MJ, Beisner J, Stange EF, Wehkamp J. Innate antimicrobial host defense in small intestinal Crohn’s disease. Int J Med Microbiol. 2010;300(1):34-40. doi: 10.1016/j.ijmm.2009.08.011
- Armbruster NS, Stange EF, Wehkamp J. In the Wnt of Paneth Cells: Immune-Epithelial Crosstalk in Small Intestinal Crohn’s Disease. Front Immunol. 2017;8:1204. doi: 10.3389/fimmu.2017.01204
- He GW, Lin L, DeMartino J, et al. Optimized human intestinal organoid model reveals interleukin-22-dependency of paneth cell formation. Cell Stem Cell. 2022;29(9):1333-1345. e6. doi: 10.1016/j.stem.2022.08.002
- Hamdan FH, Farhadipour, et al. Intestinal Stem Cells From Patients With Inflammatory Bowel Disease Retain an Epigenetic Memory of Inflammation. Cell Mol Gastroenterol Hepatol. 2026;101774. doi: 10.1016/j.jcmgh.2026.101774
- Dotti I, Mora-Buch R, Ferrer-Picón E, et al. Alterations in the epithelial stem cell compartment could contribute to permanent changes in the mucosa of patients with ulcerative colitis. Gut. 2016;66(12):2069-2079. doi: 10.1136/gutjnl-2016-312609
- Arnauts K, Sudhakar P, Verstockt S, et al. Microbiota, not host origin drives ex vivo intestinal epithelial responses. Gut Microbes. 2022;14(1):2089003. doi: 10.1080/19490976.2022.2089003
- Lee C, Hong SN, Kim ER, Chang DK, Kim YH. Epithelial Regeneration Ability of Crohn’s Disease Assessed Using Patient-Derived Intestinal Organoids. Int J Mol Sci. 2021;22(11):6013. doi: 10.3390/ijms22116013
- Schirmer M, Garner A, Vlamakis H, Xavier RJ. Microbial genes and pathways in inflammatory bowel disease. Nat Rev Microbiol. 2019;17(8):497-511. doi: 10.1038/s41579-019-0213-6
- Gevers D, Kugathasan S, Denson LA, et al. The Treatment- Naive Microbiome in New-Onset Crohn’s Disease. Cell Host Microbe. 2014;15(3):382-392. doi: 10.1016/j.chom.2014.02.005
- Rimmer P, Zhang F, Scott G, et al. The Gut Microbiome at the Onset of Inflammatory Bowel Disease: A Systematic Review and Unified Bioinformatic Synthesis. Gastroenterology. 2026;170(3):539-556. doi: 10.1053/j.gastro.2025.09.014
- Zheng L, Duan SL, Wang K. Research progress concerning the involvement of the intestinal microbiota in the occurrence and development of inflammatory bowel disease. World J Gastroenterol. 2025;31(42):113170. doi: 10.3748/wjg.v31.i42.113170
- Jalili-Firoozinezhad S, Gazzaniga FS, Calamari EL, et al. A complex human gut microbiome cultured in an anaerobic intestine-on-a-chip. Nat Biomed Eng. 2019;3(7):520-531. doi: 10.1038/s41551-019-0397-0
- Floor E, Su J, Chatterjee M, et al. Development of a Caco- 2-based intestinal mucosal model to study intestinal barrier properties and bacteria–mucus interactions. Gut Microbes. 2024;17(1):2434685. doi: 10.1080/19490976.2024.2434685
- Fofanova TY, Karandikar UC, Auchtung JM, et al. A novel system to culture human intestinal organoids under physiological oxygen content to study microbial-host interaction. PLoS One. 2024;19(7):e0300666. doi: 10.1371/journal.pone.0300666
- Han X, Mslati MA, Davies E, Chen Y, Allaire JM, Vallance BA. Creating a More Perfect Union: Modeling Intestinal Bacteria-Epithelial Interactions Using Organoids. Cell Mol Gastroenterol Hepatol. 2021;12(2):769-782. doi: 10.1016/j.jcmgh.2021.04.010
- Karve SS, Pradhan S, Ward DV, Weiss AA. Intestinal organoids model human responses to infection by commensal and Shiga toxin producing Escherichia coli. PLoS One. 2017;12(6):e0178966. doi: 10.1371/journal.pone.0178966
- Co JY, Margalef-Català M, Li X, et al. Controlling Epithelial Polarity: A Human Enteroid Model for Host-Pathogen Interactions. Cell Rep. 2019;26(9):2509-2520.e4. doi: 10.1016/j.celrep.2019.01.108
- Tan H, Chen X, Wang C, et al. Intestinal organoid technology and applications in probiotics. Crit Rev Food Sci Nutr. 2023;65(6):1055-1069. doi: 10.1080/10408398.2023.2288887
- Poletti M, Arnauts K, Ferrante M, Korcsmaros T. Organoid-based Models to Study the Role of Host-microbiota Interactions in IBD. J Crohns Colitis. 2020;15(7):1222-1235. doi: 10.1093/ecco-jcc/jjaa257
- Beaumont M, Blanc F, Cherbuy C, et al. Intestinal organoids in farm animals. Vet Res. 2021;52(1):33. doi: 10.1186/s13567-021-00909-x
- Moon C, VanDussen KL, Miyoshi H, Stappenbeck TS. Development of a primary mouse intestinal epithelial cell monolayer culture system to evaluate factors that modulate IgA transcytosis. Mucosal Immunol. 2014;7(4):818-828. doi: 10.1038/mi.2013.98
- Wang Y, DiSalvo M, Gunasekara DB, et al. Self-renewing Monolayer of Primary Colonic or Rectal Epithelial Cells. Cell Mol Gastroenterol Hepatol. 2017;4(1):165-182.e7. doi: 10.1016/j.jcmgh.2017.02.011
- VanDussen KL, Marinshaw JM, Shaikh N, et al. Development of an enhanced human gastrointestinal epithelial culture system to facilitate patient-based assays. Gut. 2014;64(6):911- 920. doi: 10.1136/gutjnl-2013-306651
- Roodsant T, Navis M, Aknouch I, et al. A Human 2D Primary Organoid-Derived Epithelial Monolayer Model to Study Host-Pathogen Interaction in the Small Intestine. Front Cell Infect Microbiol. 2020;10:272. doi: 10.3389/fcimb.2020.00272
- Sontheimer-Phelps A, Chou DB, Tovaglieri A, et al. Human Colon-on-a-Chip Enables Continuous In Vitro Analysis of Colon Mucus Layer Accumulation and Physiology. Cell Mol Gastroenterol Hepatol. 2020;9(3):507-526. doi: 10.1016/j.jcmgh.2019.11.008
- Eng SJ, Nonnecke EB, de Lorimier AJ, et al. FOXO inhibition rescues α-defensin expression in human intestinal organoids. Proc Natl Acad Sci USA. 2023;120(47):e2312453120. doi: 10.1073/pnas.2312453120
- Kim HJ, Huh D, Hamilton G, Ingber DE. Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab Chip. 2012;12(12):2165. doi: 10.1039/c2lc40074j
- Wang CM, Oberoi HS, Law D, et al. Human mesofluidic intestinal model for studying transport of drug carriers and bacteria through a live mucosal barrier. Lab Chip. 2025;25(12):2990-3004. doi: 10.1039/d4lc00774c
- Ginga NJ, Slyman R, Kim GA, et al. Perfusion System for Modification of Luminal Contents of Human Intestinal Organoids and Realtime Imaging Analysis of Microbial Populations. Micromachines. 2022;13(1):131. doi: 10.3390/mi13010131
- Williamson IA, Arnold JW, Samsa LA, et al. A High- Throughput Organoid Microinjection Platform to Study Gastrointestinal Microbiota and Luminal Physiology. Cell Mol Gastroenterol Hepatol. 2018;6(3):301-319. doi: 10.1016/j.jcmgh.2018.05.004
- Almalla A, Alzain N, Elomaa L, et al. Hydrogel-Integrated Millifluidic Systems: Advancing the Fabrication of Mucus- Producing Human Intestinal Models. Cells. 2024;13(13):1080. doi: 10.3390/cells13131080
- Lindner M, Laporte A, Block S, Elomaa L, Weinhart M. Physiological Shear Stress Enhances Differentiation, Mucus- Formation and Structural 3D Organization of Intestinal Epithelial Cells In Vitro. Cells. 2021;10(8):2062. doi: 10.3390/cells10082062
- Wang X, He L, Dong Y, et al. Mucosa-associated bacteria and metabolites in inflammatory bowel disease: from inside to insight. NPJ Biofilms Microbiomes. 2026;12(1):15. doi: 10.1038/s41522-025-00887-4
- Huang YJ, Lewis CA, Wright C, et al. Faecalibacterium prausnitzii A2-165 metabolizes host- and media-derived chemicals and induces transcriptional changes in colonic epithelium in GuMI human gut microphysiological system. Microbiome Res Rep. 2024;3(3):30. doi: 10.20517/mrr.2024.14
- Wilson SS, Tocchi A, Holly MK, Parks WC, Smith JG. A small intestinal organoid model of non-invasive enteric pathogen– epithelial cell interactions. Mucosal Immunol. 2015;8(2):352- 361. doi: 10.1038/mi.2014.72
- Sarvestani SK, Signs S, Hu B, et al. Induced organoids derived from patients with ulcerative colitis recapitulate colitic reactivity. Nat Commun. 2021;12(1):262. doi: 10.1038/s41467-020-20351-5
- Gilliland A, Chen Y, Ng I, et al. Biopsy-derived colonoid air-liquid interface monolayers reveal persistent mucosal defects in ulcerative colitis patients. Inflamm Bowel Dis. 2026;32(3):526-541. doi: 10.1093/ibd/izaf318
- Xu X, Zhang Y, Huang G, Perekatt A, Wang Y, Chen L. Advances and applications of gut organoids: modeling intestinal diseases and therapeutic development. Life Med. 2025;4(2):lnaf012. doi: 10.1093/lifemedi/lnaf012
- Day CP, Merlino G, Van Dyke T. Preclinical Mouse Cancer Models: A Maze of Opportunities and Challenges. Cell. 2015;163(1):39-53. doi: 10.1016/j.cell.2015.08.068
- Yang Z, Chen J, Xiao Y, et al. Digital Barcodes for High- Throughput Screening. Chem Bio Eng. 2024;1(1):2-12. doi: 10.1021/cbe.3c00085
- Jang KK, Hudesman D, Jones DR, et al. Tofacitinib Uptake by Patient-Derived Intestinal Organoids Predicts Individual Clinical Responsiveness. Gastroenterology. 2024;167(7):1453- 1456.e5. doi: 10.1053/j.gastro.2024.07.035
- Smabers LP, Wensink GE, Verissimo CS, et al. Patient- Derived Organoids Predict Treatment Response in Metastatic Colorectal Cancer. Clin Cancer Res. 2025;31(23):5015-5026. doi: 10.1158/1078-0432.ccr-25-1564
- Penrose HM, Sinha S, Tindle C, et al. New Approach Methodologies in Crohn’s Disease Link Molecular Disease Subtypes to Clinical Outcomes. bioRxiv. Posted online April 3, 2025. doi: 10.1101/2025.04.01.25325058
- Serra D, Mayr U, Boni A, et al. Self-organization and symmetry breaking in intestinal organoid development. Nature. 2019;569(7754):66-72. doi: 10.1038/s41586-019-1146-y
- Lukonin I, Serra D, Challet Meylan L, et al. Phenotypic landscape of intestinal organoid regeneration. Nature. 2020;586(7828):275-280. doi: 10.1038/s41586-020-2776-9
- Lavickova B, Kronabitter H, Cervera-Negueruela M, et al. Integrated Microfluidic Platform for High-Throughput Generation of Intestinal Organoids in Hydrogel Droplets. Adv Sci. 2026;13(13):2516507. doi: 10.1002/advs.202516507
- Esch MB, Mahler GJ, Stokol T, Shuler ML. Body-on-a-chip simulation with gastrointestinal tract and liver tissues suggests that ingested nanoparticles have the potential to cause liver injury. Lab Chip. 2014;14(16):3081-3092. doi: 10.1039/c4lc00371c
- Arakawa H, Sugiura S, Kawanishi T, et al. Kinetic analysis of sequential metabolism of triazolam and its extrapolation to humans using an entero-hepatic two-organ microphysiological system. Lab Chip. 2020;20(3):537-547. doi: 10.1039/c9lc00884e
- Trietsch SJ, Naumovska E, Kurek D, et al. Membrane-free culture and real-time barrier integrity assessment of perfused intestinal epithelium tubes. Nat Commun. 2017;8(1):262. doi: 10.1038/s41467-017-00259-3
- Yui S, Nakamura T, Sato T, et al. Functional engraftment of colon epithelium expanded in vitro from a single adult Lgr5+ stem cell. Nat Med. 2012;18(4):618-623. doi: 10.1038/nm.2695
- Spence JR, Mayhew CN, Rankin SA, et al. Directed differentiation of human pluripotent stem cells into intestinal tissue in vitro. Nature. 2010;470(7332):105-109. doi: 10.1038/nature09691
- Deng F, Wu Z, Zou F, Wang S, Wang X. The Hippo–YAP/TAZ Signaling Pathway in Intestinal Self-Renewal and Regeneration After Injury. Front Cell Dev Biol. 2022;10:894737. doi: 10.3389/fcell.2022.894737
- Mulero-Russe A, García AJ. Engineered Synthetic Matrices for Human Intestinal Organoid Culture and Therapeutic Delivery. Adv Mater. 2023;36(9):2307678. doi: 10.1002/adma.202307678
- Sugimoto S, Kobayashi E, Fujii M, et al. An organoid-based organ-repurposing approach to treat short bowel syndrome. Nature. 2021;592(7852):99-104. doi: 10.1038/s41586-021-03247-2
- Kasendra M, Troutt M, Broda T, et al. Intestinal organoids: roadmap to the clinic. Am J Physiol Gastrointest Liver Physiol. 2021;321(1):G1-G10. doi: 10.1152/ajpgi.00425.2020
- Kozlowski MT, Crook CJ, Ku HT. Towards organoid culture without Matrigel. Commun Biol. 2021;4(1):1387. doi: 10.1038/s42003-021-02910-8
- Ahn SJ, Lee S, Kwon D, et al. Essential Guidelines for Manufacturing and Application of Organoids. Int J Stem Cells. 2024;17(2):102-112. doi: 10.15283/ijsc24047
- Wang Y, Lin H, Zhao L, et al. Standard: Human intestinal organoids. Cell Regen. 2023;12(1):19. doi: 10.1186/s13619-023-00168-5
- Sugimoto S, Kobayashi E, Kanai T, Sato T. In Vivo Intestinal Research Using Organoid Transplantation. Keio J Med. 2022;71(4):73-81. doi: 10.2302/kjm.2022-0019-ir
- Hanna E, Rémuzat C, Auquier P, Toumi M. Advanced therapy medicinal products: current and future perspectives. J Mark Access Health Policy. 2016;4(1):31036. doi: 10.3402/jmahp.v4.31036
- Lambda Biologics. Organoids Enter the Clinic: A Milestone for Regenerative Medicine. Lambda Biologics. Published March 27, 2026. Available from: https://afs.lambda-bio. com/blog/organoids-enter-the-clinic-a-milestone-for-regenerative-medicine/ [Last accessed on April 2, 2026].
- Ouahoud S, Giugliano F, Muncan V. Monitoring Intestinal Organoid–Derived Monolayer Barrier Functions with Electric Cell–Substrate Impedance Sensing (ECIS). Bio Protoc. 2024;14(1340):e4947. doi: 10.21769/bioprotoc.4947
105. Lee H, Yang S, Lee KJ, et al. Standardization and quality assessment for human intestinal organoids. Front Cell Dev Biol. 2024;12:1383893. doi: 10.3389/fcell.2024.1383893

